ARCHIVED - Emergence of Quinolone-Resistant Shigella Dysenteriae Type 1 in Canada
Introduction
Shigella dysenteriae type 1 is an endemic human pathogen causing outbreaks of acute bacillary dysentery in high-density populations, and recent clones are resistant to a wide spectrum of antimicrobials(1). In the last 2 years, resistance to quinolone antibiotics has been reported in India and Bangladesh(2,3), presumably following the established trend of this organism to rapidly develop resistance to the current therapeutics. Moreover, this resistance profile leaves few reliable and economical therapeutic options for S. dysenteriae type 1. The incidence of shigellosis and dysentery in Canada and in the U.S. is relatively low, particularly S. dysenteriae type 1, which is rarely observed(4). From 1998 to November 2004, 110 S. dysenteriae were reported to the National Microbiology Laboratory (NML), and only three isolates were type 1. One of these was a quinolone-resistant isolate (NML stock number 04-3516) that the Alberta Provincial Laboratory of Health collected from a 56-year-old man in Calgary, Alberta in June 2004. Initially, antibiotic susceptibility testing identified isolate 04-3516 with a multidrug-resistance profile, including quinolones (ciprofloxacin and nalidixic acid). This isolate was associated with travel to India, and a concurrently isolated S. dysenteriae type 1 strain (04-3515 from a 31-year-old man in Edmonton, Alberta, with no identified travel history), had an identical antibiotic resistance profile, with the exception of a sensitivity to quinolones. We sought to identify the mechanisms of quinolone resistance encoded by isolate O4-3516, and the genetic relatedness of these Canadian S. dysenteriae type 1 isolates to those observed in south Asia.
Methods
Antibiotic susceptibility testing was completed according to Clinical and Laboratory Standards Institute guidelines for the disc diffusion method, and minimal inhibitory concentration (MIC) values were determined by broth microdilution (Sensititre, Trek Diagnostics Systems). Oligonucleotides used for polymerase chain reaction and sequencing of quinolone resistance-determining regions (QRDR) of gyrA, gyrB, parC, or parE were as previously described (see below) or designed after comparison between E. coli QRDR primers(5) and sequence data for S. dysenteriae M131649 (accession: NC_004510): parC (STPARC1(6); TDCPARC2-R: 5’ GCTCGGAATATTTCGACAACC), parE (GIL223: 5’ CACCGAACTGTTTCTTGTGG; GIL224: 5’ TGGCAATGTGCAGACCATCGG), gyrB (GIL225: 5’ CAAACTGGCAGACTGCCAGGAG; GIL226: 5’ CAGTTTGTCCGGGTTGTACTCG), and gyrA (STGYRA1 and STGYRA12 [6]). Sequence comparisons among gyrA, gyrB, parC, or parE loci encoded by S. dysenteriae M131649, 04-3515, and 04-3516 were done with ClustalW (http://www.ebi.ac.uk/clustalw/index.html) to identify mutations encoding quinolone resistance. Pulsed-field gel electrophoresis was performed using standard PulseNet protocols.
Results
To determine the mechanism(s) of ciprofloxacin resistance encoded by strain 04-3516, the QRDR of gyrA, gyrB, parC and parE were amplified and sequenced, and for comparison, the sequences of these same QRDR encoded by quinolone-sensitive strain 04-3515 were determined. In reference to S. dysenteriae M131649, the quinolone-sensitive strain 04-3515 had wild-type QRDR sequences at the four examined loci, but point mutations encoding codon changes at gyrA and parC were detected for isolate 04-3516 (Table 1). The gyrA gene of quinolone-resistant isolate 04-3516 encodes Ser83Ile and Asp87Asn-encoding substitutions. Quinolone resistance of strain 04-3516 is likely also supported by a Ser80Ile substitution in parC.
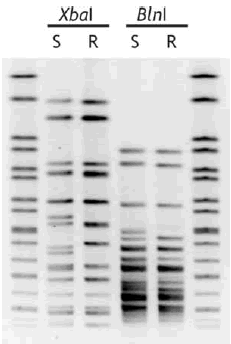
A DNA fingerprint for both the quinolone-sensitive and resistant isolate was determined by pulsed-field gel electrophoresis (PFGE) after digestion of chromosomal DNA with XbaI or BlnI (Figure 1). The XbaI banding pattern of isolate 04-3515 correlated to PFGE type A3 from sporadic Bangladeshi and Indian outbreak isolates reported by Talukder and colleagues(7). Further, 04-3516 was not represented in this data set of south Asian S. dysenteriae type 1 isolates, but of 13 DNA fragments, 12 matched the 04-3515/A3 type fragments. Analysis of plasmid content indicated identical profiles for both strains, including small molecular weight plasmids of 2, 4, and 6.3 MDa (data not shown).
Discussion
The resistance determinants encoded by S. dysenteriae type 1, such as those of other enteric pathogens, are present on integrons and conjugative plasmids, or resistance is mediated by point mutations within chromosomally encoded genes. One mechanism of quinolone resistance is by altering the drug target– DNA gyrase and DNA topoisomerase IV – each of which are type II topoisomerases. Occurrences of mutation in the gyrA and gyrB (DNA gyrase) and parC and parE (topoisomerase IV) have been reported for ciprofloxacin-resistant Escherichia coli and Salmonella spp.(8,9). Sequence analysis of the gyrA, gyrB, parC, and parE QRDR has recently been reported for Asian ciprofloxacin-resistant S. dysenteriae type 1 isolates(7,10), and we have detected identical mutations in gyrA and parC encoded by quinolone-resistant isolate 04-3516. These genetic similarities and the known travel to India associated with this isolate indicate that transmission to Canada was from this region.
Figure 1. Pulsed-field gel electrophores is a Xbal- andBinl-digested genomic DNA from ciprofloxacin sensitive (“S”, isolate 04-3515) and resistant (“R”, 04-3516) Shigella dysenteriae type 1. Salmonella Braenderup H9812 (Xbal-digested) is also run in th first and last lanes as a PFGE standard for inter-laboratory comparison
The XbaI PFGE banding patterns of both isolates 04-3515 and 04-3516 were unique to the Canadian national PulseNet database and the Center for Disease Control and Prevention database (Table 1), likely owing to the rare occurrence of this pathogen in North America. The similar banding patterns between these two isolates indicated that there are genetic differences in addition to the point mutations encoded at the gyrA and parC loci, but overall, there is considerable genetic relatedness. Notably, isolate 04-3515 had an identical pattern to S. dysenteriae type 1 strains that were previously observed in south Asia(7).While the patient identified no travel history, this genetic similarity indicates that this strain was also transmitted from south Asia to Canada. Further,similar plasmid profiles were observed between the recently isolated ciprofloxacin-resistant south Asian strains(7) and these two Canadian isolates.
Table 1. Antimicrobial susceptibility profiles and genetic analysis of Shigella dysenteriae type 1 isolated in June 2004, Alberta, Canada
| MIC (mg/L) a |
QRDR Sequencing b |
|||||||||||||||||
Strain |
Ac |
Ap |
Cip |
Nal |
St |
Su |
Tc |
Tm |
gyrA |
parC |
gyrB |
parE |
PFGE c |
Toxin d |
||||
| 04-3515 |
16 |
32 |
0.015 |
1 |
64 |
256 |
32 |
4 |
wt |
wt |
wt |
wt |
J2PX01.0004 |
stx1 |
||||
| 04-3515 |
16 |
32 |
4 |
32 |
64 |
256 |
32 |
4 |
S83L, D87N |
S80I |
wt |
wt |
J2PX01.0005 |
stx1 |
||||
a Determined by Sensititre broth microdilution, and results indicating ‘susceptible’ by disc diffusion are underlined; Ac = amoxicillin/clavulanic Acid, Ap = ampicillin, Cm = chloramphenicol, Cip = ciprofloxacin, Nal = nalidixic acid, St = streptomycin, Su = sulfamethoxoasole, Tc = tetracycline, Tm = trimethoprim b Quinolone resistance-determining regions were amplified and sequenced using primers as described in the text; ‘wt’ = wild-type in reference to S. dysenteriae M131649 (accession: NC_004510). c Pulsed-field gel electrophoresis after XbaI digestion; pattern numbers as provided by the Center for Disease Control and Prevention, Atlanta, U.S. d Toxin genotype and phenotype, determined by standard PCR and cell cytotoxicity procedures |
||||||||||||||||||
Conclusions
The introduction of foreign multi-drug resistant strains of S. dysenteriae type 1 into other global communities through travel is indicated in this study. Dominant clones of ciprofloxacin-resistant S. dysenteriae type 1 have been observed in Southeast Asia(1,11), and recent Canadian isolates are similar to Asian isolates, as indicated by gyrA QRDR sequence data and by XbaI-PFGE banding patterns. No evidence of transmission existed within the local community by the infected individuals, but the appearance of multi-drug resistant strains necessitates increased monitoring and reporting for S. dysenteriae in Canada and globally.We encourage all health centres to submit presumptive S. dysenteriae strains to provincial or national laboratories.
Acknowledgements
We acknowledge the assistance of P. Tilley and M. Louie at the Alberta Provincial Laboratory of Health for providing strains; the NML DNA Core unit for synthesis of oligonucleotides and DNA sequencing; D. Kuntz and D. Janella for performing susceptibility tests; W. Demczuk, M. Boyd, and L. Tschetter for providing information related to the strains; P. Backhouse for critical review of this manuscript; and G.B. Nair for sharing data with us.
References
-
Pazhani GP, Sarkar B, Ramamurthy T et al. Clonal multidrug-resistant Shigella dysenteriae type 1 strains associated with epidemic and sporadic dysenteries in eastern India. Antimicrob Agents Chemother 2004;48:681-84.
-
Naheed A, Kalluri P, Talukder KA et al. Fluoroquinoloneresistant Shigella dysenteriae type 1 in northeastern Bangladesh. Lancet Infect Dis 2004;4:607-08.
-
Dutta S, Dutta P, Matsushita S et al. Shigella dysenteriae serotype 1, Kolkata, India. Emerg Infect Dis 2003;9:1471-74.
-
Gupta A, Polyak CS, Bishop RD et al. Laboratory-confirmed Shigellosis in the United States, 1989-2002: Epidemiologic trends and patterns. Clin Infect Dis 2004;38:1372-77.
-
Everett MJ, Jin YF, Ricci V et al. Contributions of individual mechanisms to fluoroquinolone resistance in 36 Escherichia coli strains isolated from humans and animals. Antimicrob Agents Chemother 1996;40:2380-86.
-
Giraud E, Brisabois A, Martel JL et al. Comparative studies of mutations in animal isolates and experimental in vitro- and in vivo-selected mutants of Salmonella spp. suggest a counterselection of highly fluoroquinolone-resistant strains in the field. Antimicrob Agents Chemother 1999;43:2131-37.
-
Talukder KA, Khajanchi BK, Islam MA et al. Genetic relatedness of ciprofloxacin-resistant Shigella dysenteriae type 1 strains isolated in south Asia. J Antimicrob Chemother 2004;54:730-34.
-
Eaves DJ, Randall L, Gray DT et al. Prevalence of mutations within the quinolone resistance-determining region of gyrA, gyrB, parC, and parE and association with antibiotic resistance in quinolone-resistant Salmonella enterica. Antimicrob Agents Chemother 2004;48:4012-15.
-
Yang H, Chen S, White DG et al. Characterization of multipleantimicrobial- resistant Escherichia coli isolates from diseased chickens and swine in China. J Clin Microbiol 2004;42:3483-89.
-
Dutta S, Kawamura Y, Ezaki T et al. Alteration in the GyrA subunit of DNA gyrase and the ParC subunit of topoisomerase IV in Quinolone-resistant Shigella dysenteriae serotype 1 clinical isolates from Kolkata, India. Antimicrob Agents Chemother 2005;49:1660-61.
-
Dutta S, Ghosh A, Ghosh K et al. Newly emerged multipleantibiotic- resistant Shigella dysenteriae type 1 strains in and around Kolkata, India, are clonal. J Clin Microbiol 2003;41:5833-34.
Source: M Gilmour, PhD and T Cote, Emerging Bacterial Pathogens Program; D Woodward, BSc and L-K Ng, PhD, Bacteriology and Enteric Diseases Program, National Microbiology Laboratory, Winnipeg, Manitoba, Public Health Agency of Canada.
Page details
- Date modified: