Guidance on the Use of Quantitative Microbial Risk Assessment in Drinking Water

Download the alternative format
(PDF format, 610 KB, 40 pages)
Organization: Health Canada
Published: 2018-03-09
Document for Public Consultation
Prepared by the Federal-Provincial-Territorial
Committee on Drinking Water
Consultation period ends
May 11, 2018
Table of Contents
- Guidance on the Use of Quantitative Microbial Risk Assessment in Drinking Water
- Purpose of consultation
- Executive summary
- Assessment
- International considerations
- Part A Guidance on the use of QMRA in drinking water
- A1 Introduction and background
- A2 Determining a risk assessment approach
- A3 Sensitivity analyses: accounting for variability and uncertainty in risk assessment
- A4 Assumptions and limitations associated with risk assessments
- A41 Pathogen concentration estimates
- A42 Effectiveness of treatment barriers
- A43 Exposure analysis
- A5 Understanding risk estimates
- A6 Application of QMRA in managing water safety
- Part B Supporting information
- Part C References and acronyms
February 2018
Guidance on the Use of Quantitative Microbial Risk Assessment in Drinking Water
Purpose of consultation
The Federal-Provincial-Territorial Committee on Drinking Water (CDW) has developed this document with the intent to provide regulatory authorities and decision-makers with information on the Health Canada quantitative microbial risk assessment (QMRA) model; to describe the principles, equations, and literature values used in supporting the development of drinking water guideline values for enteric viruses and protozoa; and to provide information on the assumptions and limitations of conducting site-specific risk assessments at drinking water treatment facilities. The purpose of this consultation is to solicit comments on this guidance document.
The CDW has requested that this document be made available to the public and open for comment. Comments are appreciated, with accompanying rationale, where required. Comments can be sent to the CDW Secretariat via email at water_eau@hc-sc.gc.ca. If this is not feasible, comments may be sent by mail to the CDW Secretariat, Water and Air Quality Bureau, Health Canada, 3rd Floor, 269 Laurier Avenue West, A.L. 4903D, Ottawa, Ontario K1A 0K9. All comments must be received before May 11, 2018.
Comments received as part of this consultation will be shared with the appropriate CDW member, along with the name and affiliation of their author. Authors who do not want their name and affiliation shared with their CDW member should provide a statement to this effect along with their comments.
It should be noted that this guidance document on the use of QMRA in drinking water will be revised following evaluation of comments received, and the final document will be posted. This document should be considered as a draft for comment only.
Executive summary
Quantitative microbiological risk assessment (QMRA) is an approach that can be used by regulatory agencies and drinking water authorities to quantify the health risks from microorganisms for water sources. It follows a common approach that includes hazard identification, exposure assessment, dose-response assessment and risk characterization. QMRA examines the entire drinking water system, from the source water to the consumer, to understand the potential impacts on public health.
Health Canada has developed and uses a QMRA model to support the development of drinking water guidelines for enteric viruses and protozoa. The model can also be used as part of site-specific risk assessments at drinking water treatment facilities.
Health Canada recently completed its review of the use of QMRA in drinking water. This guidance document provides an overview of the considerations, assumptions, and limitations that are necessary for conducting site-specific risk assessments. It also describes the principles, equations, and literature values used by the Health Canada QMRA model.
During its fall 2016 meeting, the Federal-Provincial-Territorial Committee on Drinking Water reviewed the guidance document on the use of QMRA in drinking water and gave approval for this document to undergo public consultation.
Assessment
QMRA can be a very useful tool in support of water safety management decisions. A well-formulated and thoughtful QMRA can offer important information on prioritizing hazards, identifying alternative risk management priorities and options, selection of appropriate interventions, cost-benefit analysis of risk management actions and setting of health-based performance targets. It is important to remember that QMRA does not calculate actual disease outcomes, but provides a probability that disease may occur through the water system.
The intent of this document is to provide industry stakeholders, such as provincial and territorial regulatory authorities, decision-makers, water system owners, and consultants with guidance on the use of QMRA to assist in understanding microbiological risks in Canadian water systems.
International considerations
QMRA is increasingly being applied by international agencies and governments at all levels as the foundation for informed decision-making surrounding the health risks from pathogens in drinking water. The World Health Organisation, the European Commission, the Netherlands, Australia and the United States have all made important advances in QMRA validation and methodology. These agencies and governments have adopted approaches that use QMRA to inform the development of health targets and risk management for microbiological contaminants.
Part A. Guidance on the use of QMRA in drinking water
A.1 Introduction and background
The Guidelines for Canadian Drinking Water Quality encourage the adoption of a multi-barrier source-to-tap approach to produce clean, safe and reliable drinking water (Health Canada, 2013). As part of this source-to-tap approach, quantitative microbial risk assessment (QMRA) can be used. QMRA examines the entire drinking water system from pathogens in the source water, through the treatment process, to the consumer to understand the potential impact on public health. This is done following a common approach consisting of four components: hazard identification, exposure assessment, dose-response assessment and risk characterization. Following this approach, Health Canada developed a QMRA model that has been used to support the development of drinking water guideline values for enteric viruses and protozoa, and to encourage site-specific risk assessments at drinking water treatment facilities. A copy of the model can be obtained by request from water_eau@hc-sc.gc.ca.
The purpose of this document is two-fold: to provide an overview of the considerations, including the assumptions and limitations that are necessary for conducting site-specific risk assessments; and to describe the principles, equations, and literature values used by the Health Canada QMRA (HC QMRA) model. The document is divided into 2 sections. Part A provides general guidance on the use of QMRA and is intended for individuals with an interest in, or responsibility for, drinking water quality and safety. Part B provides detailed information on the HC QMRA model along with some scenarios for its application. This information is intended for those interested in better understanding and potentially applying the QMRA tool developed by HC. By capturing general QMRA considerations (Part A) and detailed HC model development information (Part B) into one document, the intention is to provide a single document that can be used in Canada to improve understanding and implementation of QMRA as part of a source-to-tap approach. This document does not provide detailed instructions on how to carry-out site specific assessments. For examples of QMRA analyses of specific drinking water supplies, the reader is referred to the Drinking Water Plant Assessments document (WRF, in preparation), or the World Health Organization's Application of Quantitative Microbial Risk Assessment for Water Safety Management (WHO, 2016).
A.2 Determining a risk assessment approach
There is a spectrum of risk assessment approaches that can be used as part of a source-to-tap, or water safety plan approach to drinking water management. They range from qualitative to quantitative approaches. The WHO publication on risk assessment (WHO, 2016) provides a good overview of the strengths and limitations of the range of risk assessment approaches, along with general advice on when and how they should be applied.
The type of risk assessment needed for any given water system should be determined on a site-specific basis. In general, the risk assessment approach used should balance the level of detail, complexity, and evidence, with the need for the use of assumptions and expert judgment, to implement an approach that is only as complicated as necessary to make decisions on risk management options (U.S. EPA, 2014; WHO, 2016).
The first step that should be undertaken when starting a risk assessment is to determine the scope of the assessment. This can be done by asking what question(s) need to be answered. Risk assessments can be initiated for a variety of reasons (U.S. EPA, 2014), including:
- to assess the potential for human risk from exposure to a known pathogen
- to determine critical control points in the drinking water system
- to determine specific treatment processes to reduce the levels of various pathogens
- to predict the consequences of various management options for reducing risk
- to identify and prioritize research needs
- to assist in epidemiological investigations
Once the scope of the problem is defined, the appropriate type of risk assessment can be determined. The assessment could be qualitative, such as a sanitary inspection, or semi-quantitative, such as the use of risk matrices. It is recommended that all risk assessments provide some level of quantitation to help guide risk managers when prioritizing tasks (WHO, 2016). For a qualitative assessment, this could be as simple as a checklist that accompanies the sanitary inspection whereby the number of 'yes' or 'no' answers determine a high, medium, or low risk component to the system.
Quantitative risk assessment approaches can range from screening assessments that use simple point estimates to full probabilistic risk assessments that include uncertainty analysis. QMRA models that use point estimates for the input variables, such as arithmetic mean values, are known as deterministic models. Probabilistic QMRA models use statistical distributions for the input variables, as opposed to single values. Defining these statistical distributions for each input variable requires more extensive data and knowledge than using a deterministic approach.
Quantitative risk assessments are also amenable to being applied as a tiered approach. For example, a screening level assessment could be used to provide guidance on whether the system is well-above, well-below, or just meeting allowable drinking water requirements. This information could then be used to help prioritize resources. In very data limited situations, resources may be better used to implement system control measures based on the results of the screening assessment, as opposed to collecting the data necessary for a more in-depth, probabilistic assessment.
A.3 Sensitivity analyses: accounting for variability and uncertainty in risk assessment
Sensitivity analyses, which include variability and uncertainty evaluations, should be incorporated in risk assessment when possible. Variability is the natural variation in the components of a system and cannot be reduced. However, it can be better characterized by collecting additional data. Variability occurs in all components of a risk assessment, including pathogen concentrations, treatment performance, and dose-response characteristics. Uncertainty, on the other hand, is a reflection of the lack of understanding or inability to accurately measure some component that affects the outcome of the risk assessment. Uncertainty can arise from numerous sources, including a lack of information on the system under evaluation; limited local data that may not be representative of the range of values expected for that system; and from the statistical distributions selected to represent the data for the system (WHO, 2016). Uncertainty can be reduced through additional characterization of model input parameters.
Variability and uncertainty are routinely included as part of in-depth probabilistic assessments (i.e., stochastic models). They are captured using statistical distributions for the input parameters in the risk assessment model, based on the available data for the system. Adequately capturing the variability and uncertainty in the input parameters for use in probabilistic models is the most common obstacle to using a stochastic approach (U.S. EPA, 2014). Variability and uncertainty can also be included in screening level assessments using point estimates (i.e. deterministic models). In deterministic models, variability and uncertainty are usually captured by using scenarios (such as best-case and worse-case assumptions). This can help risk managers understand the probable range of risks. If a screening level assessment was conducted using only the upper limit of uncertainty for each parameter, the resulting risk estimate would be unmanageably conservative and not truly representative for the population. The use of scenarios can help determine next steps, including whether a system would benefit from a more complex stochastic modelling approach to refine their risk assessment, or whether resources would be better spent mitigating risk drivers identified during the screening level assessment. Both stochastic and deterministic models can also incorporate sensitivity analysis to determine what variables have the greatest impact on the overall risk calculations (risk drivers) (U.S. EPA, 2014; WHO, 2016).
A.4 Assumptions and limitations associated with risk assessments
There are many assumptions and limitations surrounding risk assessment implementation for drinking water management. Assumptions are made by both the model developers in constructing risk assessment models, and by the analysts and managers regarding the data inputs to the risk assessment. For example, when developing a model, assumptions made by model developers include selecting the shape of the distribution to be applied to a given parameter (e.g., normal, log-normal, triangular), and determining the dose-response model that will be used for each pathogen. These assumptions are not usually modified during individual risk assessments. The assumptions included in the development of the HC QMRA model are described in Part B. For data inputs, assumptions may be needed in place of unknown or limited information, or to minimize the complexity of the assessment. In general, pathogen concentration estimates, treatment system efficacy, and exposure information are the model inputs that are subject to assumptions by risk analysts and risk managers. In order to properly interpret risk estimates, the limitations and assumptions associated with a risk assessment need to be well documented and understood.
A.4.1 Pathogen concentration estimates
Pathogen concentration estimates for a water source are limited by the amount of information available on both the uncertainty and the variability of the collected data. First, pathogen data sets tend to be small, and therefore may not fully capture the variability inherent to the system. Low pathogen densities and the episodic nature of pathogen loading add to the difficulty in capturing this variability (U.S. EPA, 2014). Also, many systems do not have any pathogen data and will need to rely on assumptions, published literature, expert judgement, or a combination of these, for these values. Second, the methods available for detecting pathogens do not recover 100 percent of the pathogens in the samples, and recovery varies between samples. This needs to be taken into account when estimating concentrations. For some pathogens, method recovery data are not routinely determined and therefore a conservative estimate of recovery may need to be applied to ensure that the risk is not underestimated. Lastly, many detection methods do not distinguish between pathogens that are capable of causing illness in humans and those that can not. This may include the detection of both viable and non-viable pathogens (e.g. using many molecular methods), or the detection of strains that are not known to cause illness in humans. Both of these situations can potentially lead to an overestimation of risk. Some authors have argued that recovery and infectivity are of similar magnitude and therefore can be assumed to cancel one another out (Regli et al., 1991; Smeets et al., 2007). However, this relationship does not have a lot of scientific justification (WHO, 2016).
Due to the limitations associated with pathogen data, they should be used in conjunction with all the other information that is available for the system when conducting a risk assessment. Other information that could be used includes information from sanitary surveys, faecal indicator monitoring, microbial source tracking research, fate and transport modeling from faecal sources, or publications from the literature on the watershed or from other watersheds with similar faecal inputs (Ashbolt et al., 2010; U.S. EPA, 2014; WHO, 2016). All of these sources of information should be considered when making decisions regarding the pathogen concentration estimates for the system, including the associated variability and uncertainty. Further information can be found in section B.2.1.
A.4.2 Effectiveness of treatment barriers
Site-specific information on treatment barrier performance will provide the highest quality risk estimate. Utilities should make every effort to gather as much information as they can on their specific system using whatever data they have available, such as design parameters or performance assessments. Many systems will not have sufficient information to fully characterize their treatment performance and will, therefore, need to make some assumptions. There are numerous types and configurations of treatment barriers used to produce safe, reliable drinking water. Most of the commonly used treatment barriers have been extensively studied, and published literature is available on how effectively they reduce microbiological contaminants. Unfortunately, the ranges in removal for the same type of barrier can span up to 6 orders of magnitude depending on numerous factors such as water quality characteristics (e.g., temperature, organic content, pre-treatment), treatment plant design and operation (e.g., geometry, media, loading rates, hydrodynamics), and climatic factors (e.g., temperature, precipitation) (WHO, 2016). This variability in barrier performance can add significant uncertainty to a risk estimate if a drinking water system needs to rely solely on literature values.
It is important to consider what level of detail is needed for the treatment system and then record all assumptions that are being made. Treatment barrier performance decisions should also consider the data that is routinely available, such as general source water quality data and operational data from the treatment plant.
A.4.3 Exposure analysis
When determining the exposure of individuals for the purposes of risk calculations, assumptions are usually made to simplify the risk assessment, as well as to apply the risk estimate to the entire population of the drinking water source. Generally, it is assumed that the route of exposure is limited to consumption of drinking water. This requires an assumption of the volume of water consumed by an individual on a daily basis. Depending on the risk assessment model, the volume of drinking water may be included as a point estimate, or as a distribution of values. Other assumptions that are commonly applied include assuming that all individuals are equally susceptible to becoming infected. Some complex risk assessments may include variables for the immune status of the population as well as the potential for secondary spread of the pathogens to others in the community. However, this level of detail is not usually available. In addition, when the environmental exposure is expected to be low, as would be the case for treated drinking water, it has been demonstrated that similar risk estimates are obtained with or without the addition of susceptibility and secondary spread variables (Soller and Eisenberg, 2008). As such, these additional variables are not included in most drinking water risk modelling.
A.5 Understanding risk estimates
Risk estimates and health targets can be expressed on different time frames and using different metrics. Microbiological risks are usually estimated for daily exposures. The daily risks are then combined into an annual risk estimate. Tolerable health risk targets are usually expressed as annual risk targets, as opposed to daily risk targets. The advantage of annual targets is they allow for some variability in the water quality. For example, infrequent higher exposures can still occur as long they are balanced by days with much lower exposures so that the combined total for the year does not exceed the annual target. When using an annual target, it is important that it be set at a level that does not allow the variability in water quality to exceed what would be tolerable over a short term event. On the other hand, a daily target could be used to avoid the risks associated with a peak occurrence (Signor and Ashbolt, 2009).
The metrics that are generally used for expressing risk include the risk of infection or illness, or a health burden estimate such as disability adjusted life years (DALYs). The Guidelines for Canadian Drinking Water Quality use an annual target risk of 1 × 10-6 DALYs per person per year. This approach was adopted from the WHO (2004). Other jurisdictions, such as The Netherlands, use the annual risk of infection as the metric for comparison to a health target. Daily targets have not yet been used to set tolerable health risks.
When interpreting risk estimates, there are numerous factors that need to be considered. First, the quality of the data that are included in the assessment needs to be understood. This includes the assumptions that were made, how they impact the risk estimates, and to what degree the variability and uncertainty has been captured during the assessment (including noting data gaps and sampling biases). Each input into a QMRA may be based, where needed, on assumptions and expert judgement, as long as the questions that need to be answered are amenable to this approach. However, if the questions that need to be answered by the risk assessment require an in-depth probabilistic assessment, then the cost of collecting the data required for the analysis needs to be weighed against the cost of making resource decisions based on assumptions.
A.6 Application of QMRA in managing water safety
QMRA can be a very useful tool in support of water safety management decisions. A well-formulated and thoughtful QMRA can offer important information on prioritizing hazards, identifying alternative risk management priorities and options, selection of appropriate interventions, cost-benefit analysis of risk management actions and setting of health-based performance targets. It is important to remember that QMRA does not calculate actual disease outcomes, but provides a probability that disease may occur through the water system (WHO, 2016). Overall, QMRA is an aid to assist in knowing your water system(s) and therefore, can provide valuable insight for risk management.Part B. Supporting information
Part B. Supporting information
B.1 HC QMRA model overview
The HC QMRA model was first developed more than 10 years ago to support the establishment of drinking water guidelines. Since its initial development, it has undergone numerous reviews and updates. One goal of these updates has been to provide a tool that can be used by stakeholders to assess, on a site-specific basis, the potential impacts of changes, in both source water quality and treatment conditions, on the estimated health risks from microbiological contamination. To ensure the model's accessibility to a large number of users, it has been developed using a widely available software platform (Microsoft Excel).
Box B1: Mathematical models for QMRA
Mathematical models have been developed by international organizations (U.S. EPA, 2005, 2006; Smeets et al., 2008; Teunis et al., 2009; Schijven et al., 2011, 2014), as well as by other groups within Canada (Jaidi et al., 2009), as a means to quantitatively assess the potential microbiological risks associated with a drinking water system. These models include the potential risks associated with bacterial, protozoan and viral pathogens.
As mentioned in part A, the first step in conducting a risk assessment is to define its scope by determining what question(s) need to be answered, and, therefore, what type of risk assessment is needed. If it is determined that a quantitative risk assessment is needed, the HC QMRA model can be used as a screening level assessment, as well as an investigative tool to estimate risk ranges based on numerous scenarios, or to conduct a sensitivity analysis. The HC QMRA model does not provide an in-depth probabilistic assessment. If this level of analysis is required, an alternate model will be needed.
The model uses source water pathogen concentrations and treatment system information entered by the user, and ingestion and dose-response information for different microbial pathogens taken from the published literature. This information is used to estimate the annual risk of infection, the annual risk of illness, and the disability adjusted life years (DALYs) per person per year associated with the input parameters. All three endpoints are displayed to allow comparison with not only the Health Canada target of 1 × 10-6 DALYs per person per year, but also with tolerable risk levels expressed using other metrics such as an annual risk of infection.
The HC QMRA model aims to provide flexibility for users to analyze their drinking water systems in multiple ways. Users can input data that reflect their current drinking water systems, or can run scenarios to look at potential impacts of changes to various aspects of their drinking water system, such as changes in source water quality or modification of the type(s) of treatment applied. Model calculations are carried out using the mean values for most parameters, as opposed to using a more conservative estimate of the value, such as the 95th percentile. This approach was taken to provide users the opportunity to be transparent about where and when safety factors are applied; these can based on site-specific knowledge where available. To capture the range of risk estimates that are possible in a water system, users should run multiple scenarios ranging from expected conditions to situations that represent conservative estimates. It is important to note that the model does not currently include risks associated with the distribution system.
The HC QMRA model can be run using site-specific data, or using multiple assumptions and expert judgement for unknown parameters. Because of this flexibility, it is important to fully document the information that is being inputted into the model, including how representative this information is believed to be for the system, to accurately interpret the risk estimates that are obtained. Sections B.2 to B.5 provide an overview of the information that needs to be entered into the HC QMRA model, considerations for obtaining this information, and the underlying assumptions and calculations being carried out to produce the disease burden estimates. Section B.6 includes example scenarios generated using model version - V15_05 Final.
B.2 Source water pathogen concentrations
The bacterial and protozoan reference pathogens used in this model are Cryptosporidium spp, Giardia lamblia, E. coli O157:H7, and Campylobacter spp. In the case of viruses, no one virus satisfied the previously mentioned criteria. Therefore, data from rotavirus, hepatitis A virus and poliovirus were used. These were selected after a careful review of candidate microorganisms.
Box B2: Reference pathogens
Although all enteric pathogens of concern to human health should be identified during a hazard assessment of a drinking water source, risk assessment models cannot consider each individual enteric pathogen. Instead, models include only specific enteric pathogens whose characteristics make them good representatives of a particular microorganism group or hazard of concern. These are referred to as reference pathogens. It is assumed that if the risk from the reference pathogens is reduced to a tolerable level, the risk from other similar pathogens will also be addressed. Ideally, a reference pathogen provides a conservative estimate of risk by representing a worst-case combination of high occurrence, high concentration and long survival time in source water, low removal and/or inactivation during treatment and a high pathogenicity for all age groups.
Cryptosporidium spp. and Giardia lamblia were selected as the reference protozoa. They are the enteric waterborne protozoa of most concern to human health in Canada. They have high prevalence rates and the potential to cause widespread disease, and pose a treatment challenge due to their resistance to chlorination. Also, dose-response models are available for both organisms.
As no single virus has all the characteristics of an ideal reference virus, the risk assessment for enteric viruses uses characteristics from several different viruses. Since rotavirus is a common cause of infection, has been associated with severe outcomes, and has an available dose-response model, the virus risk assessment uses the health effect information from rotavirus but assumes that all age groups are susceptible to infection. For drinking water treatment, the UV inactivation data from rotavirus was used, data from hepatitis A virus and poliovirus data was used for the chemical disinfectants (U.S. EPA, 1999) to reflect viruses that are more difficult to reduce during drinking water treatment. Due to limitations associated with available monitoring methods for enteric viruses, the concentration estimates in source water may also be based on total culturable enteric viruses, as opposed to only rotavirus. Norovirus was also evaluated for use as a reference virus since it has many characteristics of an ideal reference virus, including being a significant cause of viral gastroenteritis in all age groups and a published dose-response model is available (Teunis et al., 2008). However, there is much debate surrounding the model, and some suggestion that it overestimates the infectivity of noroviruses (Schmidt, 2015). As such, norovirus has not been included in the model at this time, but will be considered for future updates.
E. coli O157:H7 and Campylobacter spp. were selected as the reference bacterial pathogens for this risk model for several reasons. They are responsible for both gastrointestinal illness and more serious health outcomes, have well established dose-response models, and are reduced through treatment at a similar level to other bacterial pathogens. In addition, Campylobacter spp. have high prevalence rates. Both pathogens are also of significant concern to human health in Canada. In addition, most drinking water utilities have data on total E. coli that can be used to estimate the concentration of E. coli O157:H7 in the source water, although this value will have a high level of uncertainty.
B.2.1 Determining source water quality
Where feasible, water providers are encouraged to implement a source water monitoring program that includes monitoring for reference pathogens, to provide site-specific information on the microbiological quality of the water. Further information on sampling methods for reference pathogens can be found elsewhere (Health Canada, 2011, 2012). Pathogen monitoring information, along with the information obtained from the sanitary survey and faecal indicator monitoring, will help risk assessors provide the highest quality information to risk managers for drinking water decision making.
The goal of a monitoring program should be to sample for the organisms of interest to an extent and at a frequency that capture the most important sources of variation in microbiological source water quality. As mentioned previously, the low density of pathogens in the source waters and their episodic nature make this task difficult. Collected samples should be identified as either baseline (routine) samples or as event (incident) samples. Event samples are those collected during periods that are expected to adversely impact water quality such as flooding or storm events. Information that defines the sample as an event sample should be included so that the conditions that constituted the event are clear. This information can be used by risk assessors to help differentiate between baseline conditions and peak events, and investigate the impact that these water quality changes have on risk estimates.
Box B3: Pathogen monitoring frequencies
In the Netherlands, where a QMRA must be conducted at least every 3 years, surface waters are monitored for 4 reference pathogens: Cryptosporidium, Giardia, enteroviruses and Campylobacter. The monitoring frequency is based on the production volume of the plant and ranges from 9 to 35 samples in a 3 year period, including both routine and incident samples. All samples can be collected in a one year period to better capture variability (Schijven et al., 2011). In the United States, the Long-Term 2 Surface Water Treatment Rule required utilities to test their surface water sources for Cryptosporidium and Giardia to determine the level of treatment required. Samples were collected as close to the intake as possible, prior to treatment, and either monthly for two years or bi-weekly for one year depending on the population being served and according to a pre-approved sampling schedule.
For many drinking water systems, it may not be feasible to obtain pathogen data for some or any of the reference pathogens in the model. Therefore, expert judgement can be used in place of the missing pathogen data. Expert judgement should be based on literature values from studies of similar types of water sources, if available, and take into consideration other site-specific information such as data from faecal indicators. Although faecal indicator data are not directly linked to pathogen concentrations, the typically larger datasets of faecal indicators can provide invaluable context for risk assessment regarding the magnitude and fluctuations of faecal contamination (WHO, 2016). By combining the information on faecal contamination with knowledge of the sources, fate, and transport mechanisms for the watershed, informed estimates of missing pathogen data is possible (Medema et al., 2009). These estimates will have a high degree of uncertainty, so the scope and complexity of the risk assessment being conducted need to be amenable to this approach.
One of the limitations of missing pathogen information is the tendency to use worse-case scenario assumptions, and therefore the concentration estimates are more likely to overestimate risks. Overestimating risks could lead to costly decisions or the diversion of resources that could be better used elsewhere to protect public health. It is therefore important to include scenarios that represent the range of conditions that could be present, in addition to worse-case scenario assumptions. This should result in more informed answers to the questions laid out at the beginning of the risk assessment process. Using various scenarios, risk assessors can also determine which of the parameters have the greatest impact on the overall risk, and consequently provide some guidance on where the most benefit would be gained by reducing a parameter's uncertainty.
B.2.2 Estimating reference pathogen concentrations
Mean pathogen concentrations (per 100 L of water) and standard deviations are entered into the model and used to fit a lognormal distribution (see section B.2.3). Entering pathogen concentrations as arithmetic means and standard deviations were chosen to make the model accessible to a variety of users. When determining mean and standard deviations, risk assessors should consider how method recovery will be incorporated, and identify how data that is below the limit of detection (LOD) will be included in the calculation. The model assumes that pathogens are randomly distributed in the water, and therefore does not account for clumping of organisms that could be occurring in the water.
There are many ways in which the same data can be analysed to estimate mean pathogen concentration and standard deviation parameters. The values could be the mean concentrations for each pathogen for a given year, to show a steady-state evaluation, or they could be the mean concentrations for each individual month to assess seasonal effects. Users can also enter concentrations from the range of values that may occur for any given scenario. This might include worst case values or defined values from the distribution of values such as the 75th or the 90th percentiles. A point estimate for the pathogen concentration can also be used by entering a very small standard deviation relative to the mean pathogen concentration (e.g., if mean = 1.0 organism /100 L, set standard deviation = 0.001 organisms/100 L).
The recovery of the method is important because methods are never 100% efficient. Collecting and analysing the large volume water samples needed to detect pathogenic microorganisms requires numerous steps. Each step in the method can contribute to the loss of some of the target organism. Recovery efficiency can vary significantly between water matrices even with a standardized method. For Cryptosporidium and Giardia datasets, most samples are analysed using U.S. EPA method 1622/23/23.1. This method includes requirements to determine the recovery in the water matrix being tested. The standard methods used for the other waterborne pathogens in the model do not have the same requirements, however, where possible, it is recommended that the recovery of these methods be assessed. In the absence of recovery efficiency information, recoveries will either need to be based on other published literature or be assumed to be 100% (Schijven, 2011). For deterministic models, recovery is incorporated into risk models using a point estimate. In a stochastic model, it is usually assumed that the variability in recovery for a given method follows a beta distribution (Teunis et al., 1997; Makri et al., 2004; Pouillot et al., 2004; Signor et al., 2006; U.S. EPA, 2014). Currently, the HC model does not include recovery. It assumes that the risk assessor has accounted for recovery (either with a point estimate or a stochastic approach) prior to entering the mean and standard deviation values.
Detection methods also may not differentiate between viable human infective organisms and those that are not a human health risk, such as non-viable organisms or species that have never been associated with human infections. This could potentially overestimate the potential health impact. For example, for Giardia and Cryptosporidium detection, the routine method detects all (oo)cysts that are recovered, and it is assumed that all (oo)cysts detected in source waters are viable and equally infectious to humans, unless evidence to the contrary exists (e.g., genotyping results). For other reference pathogens, such as enteric viruses, standard methods based on cell culture detect infectious organisms but are difficult to carry out. Instead, molecular methods that do not differentiate between viable and non-viable organisms are often employed. Where possible, assessing the viability and infectivity of the reference pathogens is recommended. The HC QMRA model allows the user to modify the fraction of infectious organisms when entering their source water pathogen data, however, the default value is 1.0 (i.e. all organisms are capable of causing infection) to provide a conservative estimate in the absence of other data.
Box B4: Transforming below detection limit values
Numerous methods have been used in the literature to transform values below the detection limit into numerical values. The approach most commonly used for screening level risk assessments is to transform the below detection limit values to numerical values by assuming they are all at a fixed concentration such as at the LOD or at ½ the LOD. The method chosen will impact the concentration estimate. For example, a study using UK finished water monitoring data transformed the below detection limit results using three different methods: all LOD values were assumed to be either all zeros (minimum values), at the LOD (maximum values), or were extrapolated linearly based on the positive detections (best estimate). It was shown that the risk varied by a factor of 4 (0.6 log) from the minimum to the maximum value assumptions (Smeets et al., 2007). The impact on risk estimates has been reported to be greatest when overall pathogen concentrations are low (Smeets et al., 2007; Jaidi et al., 2009) and when datasets are small (Jaidi et al., 2009).
Ideally, to get the best estimates of source water pathogen concentrations, the volume of sample analysed would be sufficient to have an average of at least 10 organisms in the sample (Emelko et al., 2008). However, in most source waters and for most pathogens, collecting and analysing the extremely large volume of water that would be required to recover an average of 10 organisms per sample is simply not feasible. Therefore, pathogen datasets can contain a significant number of results that are close to or below the LOD. Samples below the LOD should not be included as zeros in the calculation of mean and standard deviation; although no organisms were recovered, this does not mean that the source water has a concentration of zero. If a larger volume of water was analysed to lower the LOD, or if the recovery efficiency of the method was better, it is possible that the pathogen would be detected. The HC QMRA model assumes that the concentration of microorganisms in the source water is log-normally distributed. The log-normal distribution uses the natural logarithm, and as it is not possible to take the natural logarithm of a zero value, a lognormal distribution cannot contain zero values. Further information on the lognormal distribution is provided in section B.2.4.
The LOD for microbiological methods is determined by the volume of water that is analysed and the efficiency of the recovery method. Since microorganisms are discrete particles, the theoretical limit of detection is always one organism in the volume of water analysed. Incorporating the recovery efficiency into the LOD will make the LOD slightly higher than the theoretical limit.
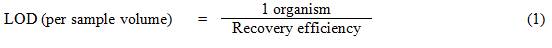
Equation 1
The Limit of detection per sample volume equals 1 organism divided by the recovery efficiency.
For example, if the sample volume is 100 L and the method used for analysis is assumed to have a recovery efficiency of 60% (expressed as a decimal fraction, i.e., 0.6), then the LOD for that sample would be calculated as 1.7 organisms per 100 L.
After deciding, and documenting how LOD data and method recovery efficiency will be addressed, mean and standard deviations can be calculated. The mean and standard deviations are entered on the Input_output worksheet of the model (see Figure B1). All pathogen concentrations, including E. coli, are entered in number of organisms per 100 L of water. The Health Canada model estimates the concentration of pathogenic E. coli, using the total E. coli data from source water(s), by assuming a default value of 3.4% of the total E. coli detected is a pathogenic strain (Martins et al., 1992). This estimate is based on raw water samples collected from a blend of Colorado River and the Northern California Water project sources. This estimate will not represent all water sources and has a high level of uncertainty. Therefore, it is not a fixed value. It can be modified in the reference worksheet of the model to best reflect the source water quality being investigated. If a drinking water system has E. coli O157:H7 data and is entering this directly, the percentage of 3.4% will need to be changed to 100%. The input parameters and the corresponding results calculated by the model will need to be recorded elsewhere as the model does not store the data for the user.
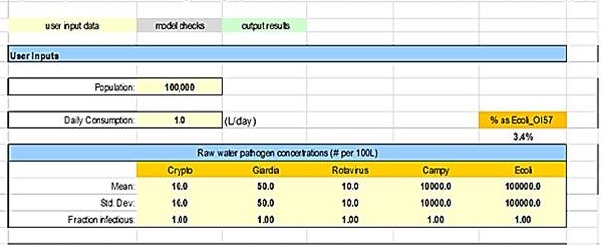
Figure B1 - Text description
A user input section of the HC QMRA model is displayed. The user has the option of entering data into the following input boxes: population; daily consumption (liters per day); and the mean, standard deviation, and fraction infectious for each of Cryptospordium, Giardia, rotavirus, Campylobacter, and E.coli. The user can also enter the value for the percent of E.coli that is E.coli O157.
Figure B1: Example of concentration and standard deviation input cells for all reference pathogens in the QMRA model (Input_Output worksheet)
B.2.3 Model calculations
Using the mean and standard deviation of the raw water pathogen concentrations entered on the Input_output worksheet, the model fits a log-normal distribution. A log-normal distribution has the shape of a normal distribution (i.e., bell shape) when you take the natural logarithm of the variable (x), in this case, the raw water pathogen concentration. The model uses the arithmetic mean (μ) and standard deviation (σ) values that were entered on the Input_output worksheet, and then estimates the mean and standard deviation of ln(x), using the following equations:
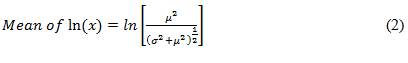
Equation 2
The mean of the natural logarithm of x (lan x) equals lan (open square bracket) the numerator is mu squared divided by the denominator of (open bracket) sigma squared plus mu squared (close bracket) to the exponent of one half (close square bracket).
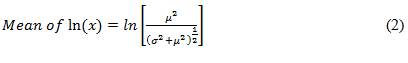
Equation 3
The standard deviation of lan x is equal to (open square bracket) lan (open bracket) sigma divided by mu (close bracket) squared plus 1 (close bracket)(close square bracket) to the exponent of one half
where,
- x = raw water pathogen concentration
- μ = mean pathogen concentration entered on Input_output worksheet
- σ = standard deviation entered on Input_output worksheet
The mean and standard deviation of ln(x) describes the shape of the lognormal distribution. The model divides the log-normal distribution curve into approximately 500 integration slices, each with an associated probability and mean concentration. This approach results in a weighted mean risk estimate. The probabilities were selected to divide the cumulative distribution function into equal segments (slices), totaling the entire area under the distribution curve. The exception is the initial portion of the curve, which is divided in smaller sections to provide better resolution at the low end of the distribution. For each integration slice, the model uses the probability for that slice and the inverse lognormal function to calculate the associated mean raw water concentration. The treated water concentration is then determined for each of the 500 slices based on the overall log-removal and log-inactivation achieved through treatment (see section B.3). The subsequent risk of infection is calculated for each slice based on the appropriate dose-response equation and is then multiplied by the probability associated with that slice of the distribution. The risk estimates are then summed to give the weighted mean risk of infection (see section B.4).
Box B5: Distributions for describing pathogen concentrations
Log-normal distributions are commonly used to describe the distribution of microorganisms in environmental samples for a couple of reasons. First, the log-normal distribution is used for skewed data. This is often the situation with raw water pathogen data where there are a large number of samples at or near the LOD and a smaller number of high concentrations. Second, it has been shown to be a reasonable fit to source water concentration data (Smeets et al., 2008; Ongerth, 2013). Other distributions have been used in the literature, such as a gamma distribution, to describe environmental pathogen data (Schijven et al., 2011, 2015). Similar to the log-normal distribution, the gamma distribution is also used for skewed data and so may also fit the source water pathogen data. In reality, no distribution will fit observed data perfectly as distributions are simple approximations to a more complicated relationship. This means several different distributions may fit the observed data equally well and the choice of distribution is determined by the researchers involved.
B.3 Determination of treatment impacts
The treatment barriers in the QMRA model are separated into 2 types: (1) physical removal methods and (2) disinfection methods. An example of the input cells for the treatment barrier information can be found in Figure B2. Physical removals for each pathogen are expressed in terms of log10 removal, whereas disinfection is expressed as log10 inactivation. The determination of log reduction values are generally based on data from surrogate parameters at full-scale treatment, or by using bench- or pilot-scale studies with laboratory-adapted strains of the pathogens of interest. These reductions are assumed to be comparable to those occurring in the treatment plant. Pathogen removal data from full-scale treatment is not usually available for log reduction calculations since pathogen concentrations naturally occurring in source water are typically low and variable.
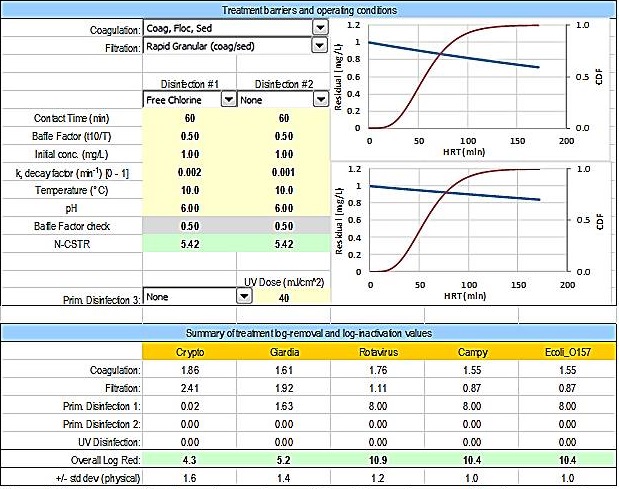
Figure B2 - Text description
This figure displays both the user input section for the treatment barriers and operating conditions, and the summary of the treatment log-removal and log-inactivation values, from the HC QMRA model. The treatment barriers and operating conditions user input section provides the following options: a drop down menu of options for coagulation; a drop down menu of options for filtration; a drop down menu of options for disinfection #1; a drop down menu of options for disinfection #2; input boxes for contact time in minutes, baffle factor in t10 over T, initial concentration in mg/L, decay factor k (per min), temperature in degrees Celsius, and pH for each of disinfection #1 and #2; a drop down menu of options for disinfection #3, with an input box for UV dose in mJ per cm2. Two graphs are also displayed in this section, representing the disinfection information input by the user for disinfection #1 and #2. Each graph displays the hydraulic residence time in minutes on the x-axis and there are two y-axes, disinfection residual in mg/L and CDF (from 0 to 1). This section also includes 2 output boxes that display a baffle factor check and the calculated N-CSTR. The second section of this figure displays the summary of treatment log-removal and log-inactivation for each of Cryptosporidium, Giardia, rotavirus, Campylobacter, and E.coli O157. The values displayed reflect the treatment barriers and operating conditions selected by the user. The overall log reduction and the standard deviation (based on the physical removal information) are also displayed.
Figure B2: Treatment barrier and operating condition information entered by the user on the Input_Output worksheet.
B.3.1 Physical removal methods
The physical removal options are separated into a coagulation step (Log RemC&S) and a filtration step (Log RemFiltr.) to provide more flexibility in representing a treatment system. The coagulation steps are the following:
- coagulation only
- coagulation and flocculation
- coagulation, flocculation and sedimentation
- none, or
- user specified
The filtration methods are:
- rapid granular (no coagulation)
- rapid granular (inline coagulation / direct filtration)
- rapid granular (with coagulation/sedimentation)
- slow sand
- membrane (micro)
- membrane (ultra)
- none, or
- user specified
For the coagulation steps, the only selection that provides log removals is the coagulation/flocculation/sedimentation option. The remaining coagulation processes do not have a particle removal step and therefore, their contributions to removals are considered part of the filtration step. Thus, for conventional treatment, both coagulation/flocculation/sedimentation and granular filtration (with coagulation/sedimentation) need to be selected to represent full conventional treatment. Also, for drinking water treatment systems that use dissolved air flotation, the removals provided for coagulation/flocculation/sedimentation are considered a reasonable estimate of this process.
The log removal data incorporated into the model for these treatment processes are based on published literature. With the exception of membrane filtration removals, the data included for each treatment stage are the weighted mean values taken from a large literature survey (Hijnen and Medema, 2007). To determine the weighted mean values for each treatment process, the authors used a weighting factor, on a scale of 1 to 5 based on the quality of the study, to calculate the weighted average log removals. For example, studies that were conducted at full-scale were given higher weight than pilot-scale studies, and studies that used pathogens as opposed to surrogates were also given greater weight. For membrane filtration, an arithmetic mean and standard deviation were calculated based on the available studies; no weighting factors were applied. The table of literature values can be found in the Treatment worksheet of the model. These data can be modified to update new pilot and full-scale research results as they become available. Relying on literature values may underestimate or overestimate the performance at a specific site. This needs to be considered by risk assessors and risk managers when making drinking water management decisions.
The option of specifying log removal/inactivation values, as opposed to using literature values, is available by selecting "user specified" and then defining the mean log-removals and standard deviation for each of the reference pathogens (Cryptosporidium, Giardia, rotavirus, E. coli, and Campylobacter). This is done in the Treatment worksheet of the model. As log reductions can vary even in well operated treatment plants, it is better to have site specific information whenever possible (Smeets et al., 2007) to be used in place of the literature values. This option is very useful for treatment plants that have carried out extensive in-house monitoring and consequently have reliable pathogen removal data demonstrating that their system performs differently than what is published in the literature. This option also provides the opportunity to investigate improvements possible through process optimization, or conversely, the impact of conditions such as suboptimal coagulation or end of filter run conditions. However, in many treatment plants, site-specific information will not be available and the drinking water system will need to rely on the pre-determined log reduction values in the model.
B.3.2 Disinfection methods
The model includes seven options:
- free chlorine
- chloramines
- ozone
- chlorine dioxide
- ultraviolet (UV) disinfection
- non, and
- user specified
The model allows for 2 stages of chemical disinfection (Log InactDisinf1 and Log InactDisinf2). To calculate the log inactivation for the chemical disinfectants (free chlorine, chloramines, ozone and chlorine dioxide), 6 parameters must be entered to describe the disinfection process (see Figure B3):
- contact time (min)
- baffle factor (T10/T)
- initial disinfectant concentration (mg/L)
- disinfectant decay factor (min-1)
- pH, and
- temperature (°C)
For UV disinfection, only the UV effective dose or fluence (mJ/cm2) needs to be entered for the treatment plant. As mentioned earlier, the log inactivation equations have generally been developed using laboratory adapted strains of pathogens. This adds some uncertainty to the calculations since environmental strains may not respond in exactly the same manner as laboratory strains.
It is important to enter data for each of the parameters that describe the disinfection process. If this information is not available for the specific treatment system, operations should be carefully examined in an effort to acquire these data. It is expected that some systems may not know their baffle factor or their disinfectant decay factor. The baffle factor can be determined accurately through tracer studies, or estimated based on the geometry of the contact chamber (i.e., inlet arrangement, baffled vs. non-baffled). The disinfectant decay factor can be assessed using jar studies or plant measurements, or can be determined through trial and error knowing the residual concentration profile through the basin. Further information can be found in section B.3.2.1.
The model uses the 6 parameters (listed above) and a continuously stirred tank reactors (N-CSTR) approach for the CT inactivation calculations of all the chemical disinfectants (details below). This approach was chosen to provide a more accurate estimate of the log inactivation being achieved in a full-scale disinfectant contact basin. This was especially important for ozone inactivation because of the fast decay rates associated with this disinfectant.
B.3.2.1 N-CSTR approach
Box B6: The N-CSTR approach
The N-CSTR approach used in the HC QMRA model is based on work published by Smeets et al. (2006). In brief, the N-CSTR approach represents the hydraulic retention time distribution using (N) theoretical continuously stirred tank reactors (CSTR) in series, based on the ratio of T10/T (i.e. the baffle factor). Baffle factors are used to account for short-circuiting in contact basins. They are the ratio of T10 to T. In a perfectly baffled system, there is no short-circuiting and the T10 and T values are almost identical. In this case, the baffle factor is equal to 1.0, also referred to as "plug-flow" conditions. In reality, disinfectant contact basins are never perfectly baffled. As mentioned previously, baffle factors can be determined accurately through tracer studies, or estimated based on the geometry of the contact chamber. Since many systems will not have conducted tracer studies, they will need to rely on estimates. Studies investigating numerous contact chamber sizes and configurations reported T10/T ratios between 0.3 and 0.7. Based on these studies, descriptions were developed to help guide water system operators in estimating a baffle factor depending on their system characteristics. These descriptions and additional information can be found in U.S. EPA (2003) and MOE (2006).
The number of N-CSTRs and the hydraulic retention time profile are first characterized, and then the distribution is divided into 1000 integration slices. The chemical inactivation is calculated for each slice using CT disinfection equations, which usually incorporate conditions of pH, temperature, and disinfectant concentration. The disinfectant concentration for each integration slice is calculated using the disinfectant decay factor. The remaining fraction of organisms is calculated for each slice and is summed over the entire basin to calculate the overall fraction of organisms remaining following the disinfection process.
For users who do not know their decay factor, it can be estimated using the following equation:
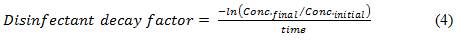
Equation 4
The disinfectant decay factor is equal to negative lan (open bracket) final concentration divided by initial concentration (close bracket) divided by time.
The initial concentration of disinfectant (Conc.initial) is the concentration (mg/L) remaining following the immediate oxidant demand. The final concentration of disinfectant (Conc.final) is the concentration (mg/L) after the contact time has elapsed, and time is the contact time (min). In general, decay factors tend to fall in the range of 0.001 to 0.2 min-1 depending on the disinfectant being applied. Alternatively, for a more conservative estimate of log inactivation, the user can enter their final disinfectant concentration as their initial concentration and set the decay factor to 0 to maintain the final disinfectant concentration through all the CSTR calculations. The user can also use trial and error to estimate the disinfectant decay factor by entering a value and reviewing the corresponding residual profile displayed to the right (see Figure B2). Once the outlet disinfectant residual matches observed operating conditions, the estimated disinfectant decay factor is reasonable.
B.3.2.2 Contact time
The value used for the contact time of the disinfectant will be dependent on the scenario that is being modelled. For a conservative inactivation estimate, the T10 time would be used. The T10 value reflects the contact time exceeded by 90% of the water in the basin. T10 is commonly used for determining CT inactivation from published CT tables. Alternatively, the mean residence time (Tmean) for the system can be entered. Tmean provides a better reflection of full-scale inactivation levels being achieved. The user can also run both scenarios (Tmean and T10) to investigate the difference in predicted treatment reductions.
B.3.3 Overall treatment reduction
Once the log-removal and log-inactivation credits are determined for the treatment processes, the overall log-reduction for each specific pathogen is calculated by adding the log removal/inactivation credits for the various treatment steps.
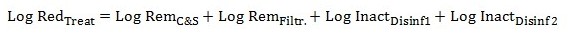
Equation 5
The overall log reduction for each specific pathogen is calculated by adding the log reductions achieved by coagulation and sedimentation, filtration, disinfection step 1 and disinfection step 2.
A summary of the log removal and inactivation values is displayed on the Input_output worksheet of the model (see Figure B2). The overall log-reduction is then used to determine the concentration of each reference pathogen in the treated drinking water.
B.4 Dose-response calculations
The goal of the dose-response calculation is to estimate the probability of infection associated with a drinking water source. To do this, the model determines the average doses of the 5 reference pathogens, calculates the probability of ingesting these doses, and finally estimates the probability of infection. This process is described in detail below.
B.4.1 Determining pathogen dose
As mentioned in section B.2, the model assumes that the raw water pathogen concentration data follows a log-normal distribution. The model divides this distribution into more than 500 integration slices to represent the total range of the distribution curve. For each of the slices, the model estimates the source water pathogen concentration and uses the overall treatment reduction to determine the treated drinking water concentration for each integration slice, as follows:
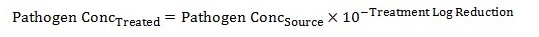
Equation 6
The pathogen concentration in treated water is calculated by taking the pathogen concentration of the source water and multiplying it by 10 to the negative exponent (total log reduction from equation 5).
The mean dose of pathogens that may be consumed by an individual is then calculated for each of the potential treated water concentrations described by the log-normal distribution, as follows:
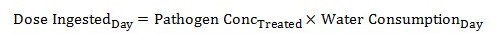
Equation 7
The pathogen concentration in treated water is calculated by taking the pathogen concentration of the source water and multiplying it by 10 to the negative exponent (total log reduction from equation 5).
The model default for average water consumption per day (Water ConsumptionDay) is 1.0 L of unboiled tap water. In a population, there will be a distribution of consumption values that are not captured by this point estimate (Statistics Canada, 2004, 2008). Data from the recent surveys of water use in Canada is included in the Reference worksheet of the model. Although the default value is 1.0 L, this value can be modified on the Input_output worksheet to reflect populations with alternative average consumptions (Figure B1).
B.4.2 Determining the probability of infection
The probability of infection is calculated using the dose-response model and parameters for each pathogen, as shown in Table B1. The exponential model has been chosen for Cryptosporidium and Giardia, whereas the beta-Poisson model is used for rotavirus, E. coli O157:H7, and Campylobacter.
| Pathogen | Dose response model | Constants | Reference |
|---|---|---|---|
| Cryptosporidium | Exponential | r = 0.018 | (Messner et al., 2001) |
| Giardia | Exponential | r = 0.01982 | (Rose and Gerba, 1991) |
| Rotavirus | Beta-Poisson | α = 0.265
β = 0.4415 |
(Haas, 1999) |
| E. coli O157 :H7Table 1 Footnote 1 | Beta-Poisson | α = 0.0571
β =2. 2183 |
(Strachan, 2005) |
| Campylobacter | Beta-Poisson | α = 0.145
β = 7.59 |
(Medema et al., 1996) |
Table 1 Footnotes
|
|||
Box B7: Dose-reponse models
Dose-response models are developed based on feeding trials, outbreak investigations, or a combination of the two. There are numerous models that could be used to describe the results from the dose-response studies, however, the models that have been shown to best describe the observed data are either the exponential model or the beta-Poisson model. Both the exponential and the beta-Poisson models are based on the single-hit theory, that is, each organism acts independently of one another and only one organism needs to survive the host-pathogen interaction in order to initiate an infection (Haas, 1999).
B.4.2.1 Exponential model (for Cryptosporidium and Giardia)
The exponential model has two main assumptions underlying its derivation. Firstly, it assumes that the number of pathogens initiating an infection is binomially distributed and that the response is the same if a single pathogen, or more than one pathogen, is responsible for the infection. Based on this assumption, the probability of at least one pathogen resulting in an infection, given a known discrete number of pathogens, can be determined using the following equation:
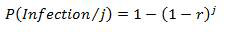
Equation 8
P(open bracket) infection divided by j (close bracket) is equal to one minus (open bracket) 1 minus r (close bracket) to the exponent j.
In this equation, j is an exact discrete number of pathogens and r is a pathogen specific constant derived from dose-response studies (Table B1).
Secondly, the exponential model assumes that the probability of ingesting an exact discrete dose of organisms (j) given an average concentration of pathogen consumed per day from drinking water (Dose Ingestedday) can be described by the following equation (i.e., a Poisson distribution):
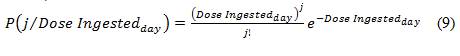
Equation 9
P(open bracket)j divided by dose ingested per day (close bracket) is equal to the numerator of (open bracket)dose ingested per day (close bracket) to the exponent j, divided by the denominator of j factorial 1, all multiplied by base e to the negative exponent of dose ingested per day.
In the HC QMRA model, discrete numbers of pathogens (j) ranging from no organisms up to a maximum of 100 organisms, in increments of 1 additional organism per dose, are used. The product of equations (8) and (9) results in the probability of infection for the parameters entered. Section B.4.3 describes how these probabilities are used.
B.4.2.2 Beta-Poisson model (for rotavirus, E.coli O157:H7, Campylobacter)
The derivation of the beta-Poisson model is similar to that of the exponential model except that the beta-Poisson model assumes that the probability of a known exact number of pathogens (j) not eliciting a response is beta-binomially distributed, as opposed to binomially distributed. Therefore, the probability of a least one pathogen resulting in an infection, out of a known number of pathogens, can be determined using the following equation:
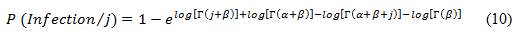
Equation 10
P(open bracket) infection divided by j (close bracket) is equal to 1 minus base e raised to the exponent log of(open square bracket) Gamma multiplied by (open bracket) j plus Beta (close bracket) (close square bracket) plus the log (open square bracket)Gamma (open bracket) alpha plus Beta (close bracket)(close square bracket) minus the log of (open square bracket) Gamma (open bracket) alpha plus Beta plus j (close bracket) (close square bracket) minus the log (open square bracket)Gamma multiplied by Beta(close square bracket).
where,
- α and β are pathogen specific parameters used to describe the ability of the pathogen to survive and initiate infection in an individual, derived from dose-response studies (Table B1), and
- Γ represents a gamma function
The second assumption of the beta-Poisson model is the same as for the exponential model: that the actual dose ingested by an individual is Poisson distributed and can be described by equation (9). The product of equations (9) and (10) results in the probability of infection for the parameters entered. An approximation to the beta-Poisson model is often used in the literature as it simplifies the equation. It was not used in this model as the assumptions for its use were not met by the reference pathogens selected. Section B.4.3 describes how the infection probabilities are used.
B.4.3 Probability of infection
The dose-response calculations in section B.4.2 determine the probability of infection for each slice of the log-normal distribution at each discrete dose between 0 and 100 organisms. This results in a large data matrix that needs to be summed by the model to provide a final probability of infection. First, for each slice of the lognormal distribution, the model calculates the probability of ingesting each discrete dose (from 0 to 100) given the mean number of organisms ingested for that slice. The probability of ingestion is then multiplied by the corresponding probability of infection and subsequently summed, to give the probability of infection for a slice of the log-normal distribution. This is done for each of the approximately 500 integration slices from the lognormal distribution. Each distribution slice is then weighted using the probability of the pathogen concentration occurring from the log-normal distribution. The weighted probabilities of infection are then summed to give the overall probability of infection per day (Pinfection,day). This value is displayed on the Input_Output worksheet (see Figure B3).
In an effort to reduce model running times, the limit of 100 organisms was applied as a compromise between realistic Canadian drinking water source contamination and treatment scenarios. It is expected that the concentration of any given pathogen in drinking water in Canada should be well below 100 organisms as an average dose. Since the discrete dose upper limit was set at 100 organisms, this model cannot be used to examine scenarios where the average ingested dose is greater than this value. For example, if a user enters data that represents a drinking water source where there is no treatment and the source water is highly contaminated such that the average pathogen dose ingested is above 100, the model will incorrectly estimate very low probabilities of infection and illness because the concentration of pathogens is outside of the analysis range. Such a scenario raises a flag in the model to alert the user.
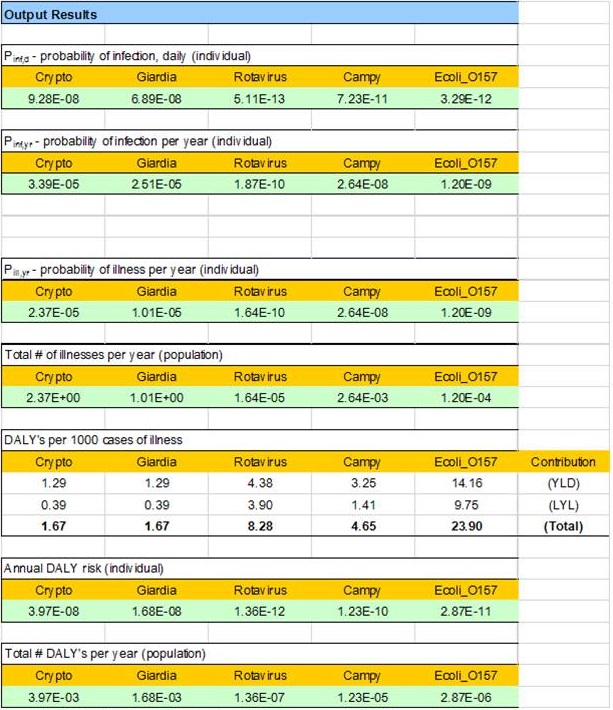
Figure B3 - Text description
The output results for the HC QMRA model are displayed. This includes the following calculated values for each of Cryptospordium, Giardia, rotavirus, Campylobacter, and E.coli: the daily probability of infection for an individual; the probability of infection per year for an individual; the probability of illness per year for an individual; the total number of illnesses per year in the population; the DALY's per 1000 cases of illness, presented as the YLD, the LYL, and the total contribution (respectively); the annual DALY risk for an individual; and the total number of DALY's per year in the population.
Figure B3: Health impact-related output values determined from user inputs.
The following equation is used to calculate the probability of 1 or more infections per year, given a daily risk of infection (Pinfection,day) (WHO, 2016):

Equation 11
The probability of one or more infections per year is calculated by subtracting from 1, the result of the term (1 minus the probability of infection) to the exponent 365.
This equation assumes that the risk of infection is the same every day for the entire year and that there is no resistance or immunity acquired in the population. In reality, the risk of infection will vary from day to day based on changes in source water quality, treatment efficiency and volume of water consumed. In addition, previous infection with some enteric pathogens provides protective immunity from subsequent infections. However, the purpose of the HC QMRA model is not to predict the number of infections or illnesses in a population, but to provide a probability that disease may occur based on the source water quality and treatment system information. Since the expected annual risk of becoming ill from drinking water is very low, the above assumptions are reasonable.
B.5 Estimating health impacts
The final step in the HC QMRA model is to determine the burden of disease, expressed in DALYs, associated with the input scenario for each of the reference pathogens. DALYs are used in this risk assessment model as a common metric to compare illnesses with different health endpoints. It is also used to allow comparisons to an established health target, in this case, the reference level of 10-6 DALYs per person per year. The calculated burden of disease values are displayed on the Input_Output worksheet (see Figure B3).
B.5.1 Determining probability of illness
To determine the disease burden for each reference pathogen, it is first necessary to calculate the probability of becoming ill, given that infection has occurred, as not all infections lead to illness. Some infected individuals may clear the infection without ever having developed any symptoms. Others might be asymptomatic carriers of the infection. These individuals also have no symptoms, but they do continue to shed the pathogen in their feces.
The probability of illness given infection (Pill/inf) varies with each reference pathogen. The values used by the model for Pill/inf are based on the published literature and are given in Table B2. They are also found in the Reference worksheet of the model. For E. coli O157:H7, the dose-response model estimates the probability of illness, so Pill/inf is 1.0 for the risk calculations. For Campylobacter, current studies do not provide a consistent value for the Pill/inf. The relationship seems to be dose dependent, with some doses showing a Pill/inf of 1.0. However, further studies are needed. In the interim, the Pill/inf has also been set at 1.0 as a conservative estimate.
| Pathogen | P(ill/inf) | Reference |
|---|---|---|
| Cryptosporidium | 0.70 | Casmen et al., 2000 |
| Giardia | 0.40 | Nash et al., 1987 |
| Rotavirus | 0.88 | Havelaar and Melse, 2003 |
| E. coli O157:H7 | 1.0 | Strachan, 2005 |
| Campylobacter | 1.0 | Assume all infections lead to illness |
The probability of illness per year for an individual (Pillness,yr) is calculated using the following equation. This calculation is carried out for each of the 5 reference pathogens.
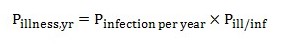
Equation 12
La probabilité d'au moins une infection par année est calculée par soustrayant de 1, le résultat du terme (1 mois la probabilité d'une infection) à l'exposant 365.
The total number of illnesses in a population can be calculated for each pathogen by multiplying the annual risk of illness times the population.
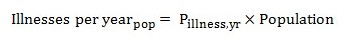
Equation 13
The total number of illnesses in a year in a population is calculated by multiplying the probability of illness in a year times the population
B.5.2 Calculating DALYs
Although estimating the probability of illness per person per year is informative, it does not provide a clear indication of the health burden associated with the drinking water source as the magnitude of the health impacts vary for each reference pathogen. As mentioned previously, DALYs are used as a common metric to compare illnesses with different health endpoints. DALY's include life-years-lost (LYL) to calculate the impact of premature death due to illness, as well as years lived with a disability (YLD) to calculate the morbidity associated with an illness.
B.5.2.1 Calculating LYL
Since premature death eliminates potential years of healthy living, the LYL is calculated as the difference between the age at death and the full life expectancy for the population, multiplied by the severity weight associated with loss of life and the fraction of ill individuals who experience the outcome (referred to as the outcome fraction).
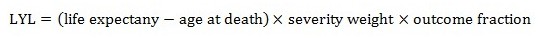
Equation 14
The LYL is calculated by subtracting the age at death from the life expectancy, then multiplying the result by the severity weight then by the outcome fraction.
The model uses the combined life expectancy (i.e. the average of male and female life expectancies - 80.88 years), as the reference pathogens do not have gender specific health outcomes. For Cryptosporidium, Giardia, rotavirus, and E.coli O157, the weighted median age (38.98 years) is used as the age at death. This assumes that there is no difference in fatality rates between the age categories. For Campylobacter, death primarily occurs in the elderly population. Therefore, the age at death is assumed to be the median age of the eldest population category (72.94 years). Complete tables of life expectancy and age values for the Canadian population can be found in the Reference worksheet of the model. In all instances, a severity weight of 1.0 is assigned for loss of life. Table B3 provides the severity weights and outcome fraction information for each reference pathogen. The outcome fractions for death are assumed to be 1 in 100,000 for Cryptosporidium and Giardia. Rotavirus and Campylobacter are assumed to have a case fatality ratio of 1 in 10,000. For E.coli O157:H7, the risk of death is higher, at 1 in 4,000 (Havelaar and Melse, 2003).
B.5.2.2 Calculating YLD
To calculate the years lived with a disability (YLD), the outcome fraction is multiplied by the severity weight (Table B3) and the duration of the illness (Table B4) for each illness outcome that is attributed to the pathogen. These products are then summed to give the YLD per case of illness.

Equation 15
The YLD per case is calculated by taking the sum of the outcome fraction times the product of the duration of illness multiplied by the severity weight
| Illness outcome | Severity weightTable 3 Footnote e | Outcome fractions | ||||
|---|---|---|---|---|---|---|
| Crypto-sporidiumTable 3 Footnote c | GiardiaTable 3 Footnote a | RotavirusTable 3 Footnote a | CampylobacterTable 3 Footnote b | E.coli O157Table 3 Footnote b | ||
| Mild diarrhea | 0.067 | 0.99999 | 0.99999 | 0.5 | 1 | 0.53 |
| Bloody diarrhea | 0.39 | - | - | 0.5 | 0.06 | 0.47 |
| Guillain-Barré syndrome (GBS) | - | - | - | - | 0.0002 | - |
| clinical and residual | ||||||
| GBS; residual | - | - | - | - | 0.0002 | - |
| Reactive arthritis | - | - | - | - | 0.023 | - |
| hemolytic uremic syndrome (HUS) | 0.93 | - | - | - | - | 0.01 |
| end-stage renal disease (ESRD) | 0.95Table 3 Footnote d | - | - | - | - | 0.00118 |
| Death (GBS) | 1 | - | - | - | 4.6E-06 | - |
| (1 in 217,000) | ||||||
| Death | 1 | 0.00001 | 0.00001 | 0.0001 | 0.0001 | 0.00025 |
| (1 in 100,000) | (1 in 100,000) | (1 in 10,000) | (1 in 10,000) | (1 in 4,000) | ||
Table 3 Footnotes
|
||||||
| Illness Outcome | Cryptosporidium | Giardia | Rotavirus | Campylobacter | E.coli O157 |
|---|---|---|---|---|---|
| mild diarrhea | 0.01918 | 0.01918 | 0.01918 | 0.01397 | 0.00932 |
| (7 days) | (7 days) | (7 days) | (5.1 days) | (3.4 days) | |
| serious diarrhea (i.e., bloody) | - | - | 0.01918 | 0.023 | 0.01534 |
| (7 days) | (8.4 days) | (5.6 days) | |||
| HUS | - | - | - | - | 0.0575 |
| (21 days) | |||||
| ESRD | - | - | - | - | 9.35 |
| clinical (GBS) | - | - | - | 0.29Table 4 Footnote a | - |
| residual (GBS) | 5.8Table 4 Footnote a | ||||
| reactive arthritis | 0.115 | ||||
| (42 days) | |||||
| death (GBS) | - | - | - | (e* - adeath) | - |
| death | (e* - adeath) | (e* - adeath) | (e* - adeath) | (e* - adeath) | (e* - adeath) |
Table 4 Footnotes
|
|||||
B.5.2.3 Total DALYs
The total DALYs per case of illness is the sum of the YLD and the LYL. In the case of a waterborne pathogen causing mild gastrointestinal illness and having a low-case fatality ratio, it is common to see the disease burden expressed in terms of DALYs per 1000 cases to illustrate the health impact more clearly. Figure B3 shows the estimated DALY's per case of illness for each of the reference pathogens including the contribution from morbidity (YLD) and mortality (LYL). With the exception of rotavirus, these DALY values are very similar to comparable values reported in the literature (Gibney et al., 2014). The DALYs per 1000 cases calculated using this model for rotavirus are approximately three times greater than the study in the literature due to differing assumptions made surrounding the susceptible population and the duration of illness.
The DALYs per person per year is then calculated by multiplying the probability of illness per person per year by the DALYs per case of illness for each pathogen. These values can then be compared to the health target of 1 × 10-6 DALYs per person per year as recommended in the Guidelines for Canadian Drinking Water Quality. The DALYs per person per year (annual DALY risk [individual]) are displayed on the Input_output worksheet of the model (Figure B3). The model also displays the total DALYs per year in the population based on the population that is added by the user on the Input_Output worksheet.
B.6 Using the model - an example case study
The following sections present a brief example of how the QMRA model can be used. This example is not intended to provide a step-by-step procedure for conducting a risk assessment using the model. It is a simplified approach, included for illustrative purposes only, to show how the QMRA model can help make risk management decisions in a drinking water system. Other examples of risk assessments can be found elsewhere (U.S. EPA, 2005, 2006; Medema et al., 2009; Schijven et al., 2011; WHO, 2016).
B.6.1 Scope of the risk assessment
The municipal drinking water system in this fictional case study is supplied by a surface water treatment plant that draws raw water from a large river. The municipality has been conducting pathogen monitoring for two years on a monthly basis (details below). The samples were collected on a pre-set schedule. The sampling conditions (i.e., baseline or incident) were not recorded. With their dataset of pathogen monitoring results, and information on the treatment processes in place, the municipality wants to conduct a screening level risk assessment to determine where they fall in relation to the tolerable risk level of 1 × 10-6 DALYs per person per year (baseline risk estimate). In addition, the municipality would like to investigate how that risk level could be impacted by challenging water conditions, altering a disinfection barrier under baseline and challenging conditions, as well as the effectiveness of an alternative physical removal method. These analyses will be part of the evidence used to help support upcoming decisions surrounding modifications to, and future expansions of, the current treatment system.
The system will use the pathogen monitoring data it has collected to estimate mean and standard deviations for each of the reference pathogens. As there is no site-specific information on pathogen log reductions, the literature values from the model will be used. The route of exposure is assumed to be only through the ingestion of unboiled tap water with an average per capita consumption of 1.0 litre of unboiled tap water per day.
B.6.2 Baseline risk estimate
The watershed surrounding the river is largely wilderness, and the river generally has low turbidity (3-5 NTU), high colour (35 true colour units) and dissolved oxygen content (6.5 mg/L). There are only a few small communities upstream of the city, with minimal wastewater discharges. Large numbers of waterfowl (Canada geese, gulls and shorebirds) can be found on the river during migration, and some overwinter in areas that do not freeze completely. A few major tributaries drain agricultural areas, which may contribute both nutrients and pathogens from animal waste to the river. Monitoring data for various pathogens in raw water were collected over two years. The data sets for all pathogens, with the exception of E.coli, contained samples that were below the LOD for the method. All below LOD samples were included in the concentration estimates assuming the concentrations were at the LOD. They were also corrected for recovery based on information received from the laboratory (when available).
The pathogen concentrations did not have clear seasonal trends, and since the samples where not identified as baseline or incident samples, the data from the 2 years of monitoring (24 samples) was combined into a single mean and standard deviation estimate (Table B5). These estimates, although they do capture variability in the pathogen concentrations, will still have uncertainty arising from the limited number of samples taken, and from limitations in pathogen detection methods, including determining whether the pathogens are human infectious. However, for the purposes of the scenarios being investigated by the utility, the uncertainty in these values will not be further explored.
| Pathogen | Cryptosporidium
(no./100 L) |
Giardia
(no./100 L) |
Rotavirus
(no./100 L) |
Campylobacter
(cfu/100 mL) |
E.coli
(cfu/100 mL) |
|---|---|---|---|---|---|
| Mean | 8.0 | 34.0 | 56.0 | 10.0 | 55.0 |
| Standard deviation | 12.0 | 72.0 | 62.0 | 22.0 | 100.0 |
cfu = colony-forming unit |
|||||
The water treatment plant has a conventional treatment process that includes: coarse screening, coagulation, flocculation, sedimentation, dual-media filtration, chlorine disinfection (primary disinfection), pH adjustment and chloramination (secondary disinfection). It is assumed that there is no further treatment or disinfection of the tap water prior to consumption, and that the quality of the treated water does not deteriorate in the distribution system.
Physical removal performance for the treatment process is not known, so the weighted mean average efficiencies for coagulation/sedimentation and filtration - rapid granular (with coag./sed.) from the QMRA model will be used. The values for these two steps are added to give the overall log removal. Using the values from the QMRA model will add uncertainty to the estimated risks, as site-specific log removals could be orders of magnitude different from the average values included in the model. However, as a screening assessment, the utility decided that average values from the literature were acceptable for the scenarios they wanted to investigate.
For primary disinfection, the chlorine residual immediately following the initial demand is 0.50 mg/L followed by a 60-minute contact time (pH=6.0, temperature=10°C). The contact time is based on mean detention time, rather than the T10 value, as this assessment is aimed at estimating the mean reduction through the treatment process. The contact basins have been properly constructed to minimize short circuiting and the baffle factor has been determined to be 0.65. The disinfectant decay factor for chlorine in this system is 0.002.
Using the QMRA model and the log reductions shown in Table B6, the mean burden of disease estimates for Cryptosporidium, Giardia, rotavirus, Campylobacter, and E.coli O157:H7 (in DALYs/person per year) were 3.27 × 10−8 , 3.58 × 10−8, 7.63 × 10−12, 1.23 × 10−10, and 1.58 × 10−10, respectively. The distribution of the estimates is shown in Figure B4. Levels of illness in this range would not reasonably be detected and, with the exception of the extreme tail of giardiasis, are well below the reference level of risk of 10−6 DALYs/person per year.
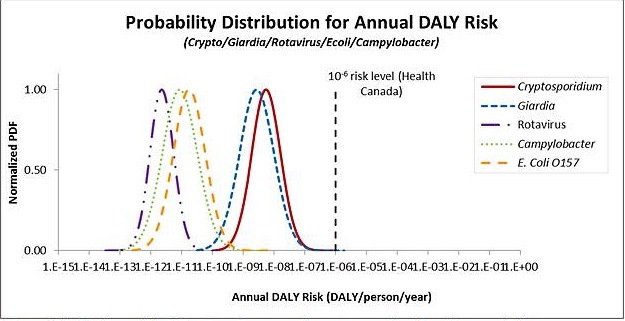
Figure B4 - Text description
A graph showing the distribution of the annual burden of illness estimates from the consumption of drinking water produced using baseline treatment conditions. The distribution of the burden of illness estimates are shown for each of Cryptosporidium, Giardia, rotavirus, Campylobacter, and E. coli O157:H7. The x-axis of the graph is the risk expressed as DALYs per person per year. It is presented on a logarithmic scale and the axis values range from 10-15 to 1. The y-axis of the graph is the normalized probability distribution function. It is presented on a linear scale and the axis values range from 0 to 1. The reference level of risk of 10-6 DALYs per person per year is illustrated on the graph as a dotted vertical line. The annual illness estimates for each pathogen are normally distributed. The distribution curves extend approximately from: slightly less than 10-10 to 10-7 for Cryptosporidium; slightly more than 10-11 to 10-6 for Giardia; slightly less than 10-14 to slightly more than 10-11 for rotavirus; 10-13 to slightly more than 10-9 for Campylobacter, and slightly more than 10-13 to almost 10-8 for E. coli O157:H7. The peak of each probability distribution is normalized to 1.
Figure B4: Estimated risk for the reference pathogens assuming baseline conditions
B.6.3 Treatment barrier modifications
Using the same pathogen data (Table B5), the QMRA model was used to investigate how modifying the treatment barriers might affect the quality of the water being produced. The QMRA model calculates risk estimates based on annual exposure, therefore the model assumes that the values used occur in the system each day for the entire year. This value is used for comparison to tolerable health risk targets. The model also displays the daily individual risk of infection, if needed. Although a constant daily risk is unlikely to occur, as water quality and treatment performance will vary throughout the year, this assumption allows decision makers to compare treatment scenarios (i.e., investigate relative risks). The fictional system wanted to investigate the impact of the following potential treatment variations as separate scenarios: (1) effectiveness of chlorine disinfection during challenging conditions (i.e., cold water temperatures and reduced contact time), (2) implementation of ozone as an alternative disinfectant to chlorine and (3) effectiveness of a membrane filtration plant in comparison to conventional treatment.
For the first scenario, the QMRA model inputs were set to represent challenging disinfection conditions. The water temperatures in the system fluctuate from 1°C to 21°C throughout the year. The system relies on chlorine for disinfection of pathogens, and the inactivation rates for this chemical are temperature dependent. Therefore, the QMRA model was run using the same parameters as the baseline estimate, with the following modifications: the temperature of the water was set to 1°C and the contact time used was the T10 value of 20 minutes to represent a conservative estimate for chlorine inactivation. The log reductions under these conditions are show in Table B6. Since chlorine inactivation is ineffective against Cryptosporidium, the challenging disinfection conditions have no impact on the reduction of this pathogen through treatment. Conversely, the bacterial pathogens are highly susceptible to inactivation by chlorine. Therefore, even using the conservative estimates, the maximum inactivation applied by the model (>8.0 log) is still achieved. The greatest impact on estimated risk is from Giardia (2.95 × 10-7 DALYs/person year) and rotavirus (3.84 × 10-10 DALYs/person year). The estimated risk from Giardia increased by approximately 1 log and the annual estimated risk for rotavirus increased by approximately 1.7 log. Both are still below the annual target, although Giardia is now within one order of magnitude and further investigation to determine the uncertainty around the log reductions being achieved may be warranted.
| Process (log10) | Cryptosporidium | Giardia | Rotavirus | Campylo-bacter | E. coli |
|---|---|---|---|---|---|
| Baseline conditions | |||||
| Coagulation/
sedimentation |
1.86 | 1.61 | 1.76 | 1.55 | 1.55 |
| Filtration - Rapid granular (with coag/sed) | 2.41 | 1.92 | 1.11 | 0.87 | 0.87 |
| Chlorine inactivation
(10°C, 60 min) |
0.0 | 1.15 | > 8.0 | > 8.0 | > 8.0 |
| Modified treatment | |||||
| Chlorine inactivation
(1°C, 20 min) |
0.0 | 0.23 | 6.3 | > 8.0 | > 8.0 |
| Ozone inactivation
(10°C, 20 min) |
0.25 | 4.0 | 4.0 | 8.0 | 8.0 |
| Ozone inactivation
(1°C, 20 min) |
0.10 | 3.79 | 4.0 | 8.0 | 8.0 |
| Membrane filter (microfiltration) | 6.13 | 6.62 | 1.10 | 4.60 | 4.60 |
For the second scenario, the fictional utility wanted to investigate the use of an alternative primary disinfectant, in this case, ozone. For this scenario, it was assumed that coagulation/sedimentation and filtration were unchanged, however, the primary disinfectant being added was ozone, as opposed to chlorine. Both baseline and challenging water conditions were considered for comparison. The ozone concentration immediately following the initial demand was projected to be 0.50 mg/L followed by a 20-minute contact time [pH = 6.0, temperature = 10°C (baseline) or 1°C (challenging)]. Since ozone is known to react very quickly in water, the contact time was set at the T10 value for both scenarios. The baffle factor was still 0.65. The disinfectant decay factor for ozone in this system was assumed to be 0.2.
The mean burden of disease estimates for Cryptosporidium, Giardia, rotavirus, Campylobacter, and E.coli O157:H7 (in DALYs/person per year) were 1.89 × 10−8, 5.02 × 10−11, 7.63× 10−8, 1.26 × 10−10, and 1.61 × 10−10, respectively, for the baseline scenario. The distribution of the estimates is shown in Figure B5. For the challenging water quality conditions, the mean burden of disease estimates for Cryptosporidium and Giardia were the only estimates that changed, with the estimated risk increasing by approximately 0.1 and 0.2 log, respectively. The utility then used the estimated ozone log reduction values for cold water to compare to the baseline chlorine disinfection estimates. This comparison showed that the estimated risk from Cryptosporidium, Campylobacter, and E.coli O157:H7 remained approximately the same. A very low level of inactivation of Cryptosporidium occurred when using ozone (0.1 log), compared to none with chlorine. The maximum inactivation allotted by the model was achieved for both of the reference bacteria, as they are highly susceptible to both disinfectants. Giardia, on the other hand, were more easily inactivated with ozone and so the estimated risk decreased by approximately 3 log. Conversely, the estimated risk from rotavirus increased by approximately 4 log, however, the model caps the log inactivation using ozone at 4 log less than for chlorine based on available published studies making direct comparison difficult. All of the estimated risks were below the annual health target.
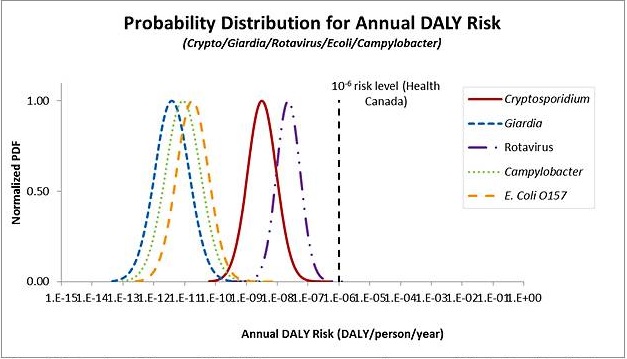
Figure B$1 - Text description
A graph showing the distribution of the annual burden of illness estimates from the consumption of drinking water produced using ozone disinfection under baseline treatment conditions. The distribution of the burden of illness estimates are shown for each of Cryptosporidium, Giardia, rotavirus, Campylobacter, and E. coli O157:H7. The x-axis of the graph is risk in DALYs per person per year. It is presented on a logarithmic scale and the axis values range from 10-15 to 1. The y-axis of the graph is the normalized probability distribution function. It is presented on a linear scale and the axis values range from 0 to 1. The reference level of risk of 10-6 DALYs per person per year is illustrated on the graph as a dotted vertical line. The annual illness estimates for each pathogen are normally distributed. The distribution curves extend approximately from: slightly less than 10-10 to almost 10-6 for Cryptosporidium; slightly less than 10-13 to 10-9 for Giardia; slightly more than 10-10 to slightly less than 10-6 for rotavirus; 10-13 to slightly more than 10-9 for Campylobacter, and slightly more than 10-13 to almost 10-8 for E. coli O157:H7. The peak of each probability distribution is normalized to 1.
Figure B5: Burden of disease in DALYs/person per year using ozone disinfection (baseline) (scenario 2).
In the third scenario, the fictional utility wanted to investigate the use of an alternative physical removal process as part of a future water treatment plant expansion project. For this scenario, it was assumed that chlorine disinfection was occurring under the conditions of the baseline estimates (10°C, 60 min) for the current plant. The alternative physical removal process was assumed to be microfiltration (Table B6). The mean burden of disease estimates for Cryptosporidium, Giardia, rotavirus, Campylobacter, and E.coli O157:H7 (in DALYs/person per year) were 4.59 × 10−10, 1.56 × 10−10, 4.50 × 10−10, 8.29 × 10−13 and 1. 06 × 10−12, respectively. The distribution of the estimates is shown in Figure B6. With the exception of rotavirus, the microfiltration scenario resulted in estimated risks equivalent to, or lower than, the conventional treatment plant. Rotavirus risks were greater but still well below the annual health target.
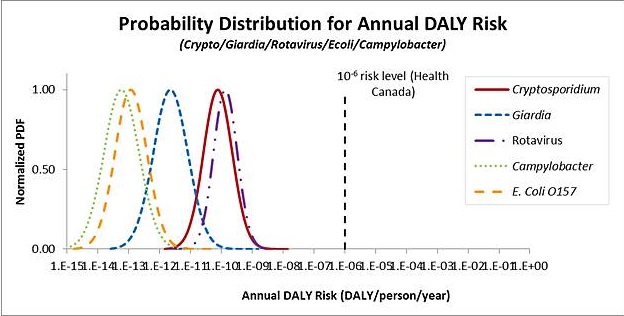
Figure B6 - Text description
A graph showing the distribution of the annual burden of illness estimates from the consumption of drinking water produced using membrane filtration as per scenario 3. The distribution of the burden of illness estimates are shown for each of Cryptosporidium, Giardia, rotavirus, Campylobacter, and E. coli O157:H7. The x-axis of the graph is the annual DALY risk in DALYs per person per year. It is presented on a logarithmic scale and the axis values range from 10-15 to 1. The y-axis of the graph is the normalized probability distribution function. It is presented on a linear scale and the axis values range from 0 to 1. The reference level of risk of 10-6 DALYs per person per year is illustrated on the graph as a dotted vertical line. The annual illness estimates for each pathogen are normally distributed. The distribution curves extend approximately from: slightly more than 10-12 to slightly more than 10-8 for Cryptosporidium; slightly more than 10-14 to slightly more than 10-9 for Giardia; slightly more than 10-12 to slightly more than 10-9 for rotavirus; 10-9 for Campylobacter, and slightly more than 10-15 to almost 10-10 for E. coli O157:H7. The peak of each probability distribution is normalized to 1.
Figure B6: Burden of disease in DALYs/person per year using membrane filtration (microfiltration) (scenario 3).
B.6.4 Interpreting risk estimates
Using the QMRA scenarios in this case study, the drinking water treatment plant appears to be producing drinking water of an acceptable microbiological quality, assuming that the conditions entered into the QMRA model are reflective of the drinking water system. From the treatment reductions applied in the model (Table B6), it is clear that the log reductions attributable to the physical removal processes are essential in controlling the risk from Cryptosporidium. Chlorine and ozone under the conditions entered offer negligible inactivation. Giardia, on the other hand, is reduced by both physical removal and disinfection. However, under challenging disinfection conditions, such as chlorine disinfection in cold water, the risk from Giardia may be approaching the target level based on the data entered in the model. This result should prompt this fictional utility to conduct some further investigation to better characterize the probable risk from Giardia to determine whether the system is meeting their health targets under all water conditions. For the bacterial and viral pathogens (Campylobacter, E.coli O157:H7 and rotavirus), disinfection is the main barrier for reducing these organisms to acceptable levels. Using the results from scenario 2, it was noted that viral pathogens are more resistant to disinfection than bacterial pathogens. The challenging water conditions with chlorine disinfection led to a probability of risk that was almost 2 orders of magnitude greater than the baseline conditions, albeit still well below the tolerable risk level. However, ozone disinfection appeared to be unaffected by the lower temperature used for this case study. Ozone was also more effective for controlling the risks from Giardia. Scenario 3 for this fictional utility, investigating the use of microfiltration as opposed to conventional filtration, showed that under the conditions of the model, microfiltration could be used to produce acceptable water quality.
The above scenarios show the relative impact on pathogen reductions based on the input parameters and the assumptions used. Understanding the assumptions made in the model is key to properly interpreting the results of the scenarios. For example, the scenario for this fictional utility assumes that the utility's physical removal processes are achieving the weighted mean log removals reported in the literature. This may over- or underestimate the log reductions being achieved by several orders of magnitude. Although this is an example of an assumption that could have a large impact on the scenario results, other parameters have less variability and therefore would have very little impact. Understanding these impacts is part of interpreting the results from the scenarios investigated. The above scenarios would be used as a starting point for the fictional utility to run further analyses that explore the range of impacts that could be expected by altering the treatment performance. The utility may also want to explore further the variability in the pathogen concentrations to determine the impact on the estimated risks. The relative risk levels could then be used to help support decisions on alternative treatment processes, or to determine what input parameters would benefit from further exploration into their site-specific variability to reduce some of the uncertainty in the results.
Part C. References and acronyms
C.1 References
Armstrong, T.W. and Haas, C.N. (2008). Legionnaires' disease: Evaluation of a quantitative microbial risk assessment model. J. Water Health, 6(2): 149-166.
Ashbolt, N.J., Schoen, M.E., Soller, J.A. and Roser, D.J. (2010). Predicting pathogen risks to aid beach management: The real value of quantitative microbial risk assessment (QMRA). Water Res., 44(16): 4692-4703.
Casman, E.A., Fischhoff, B., Palmgren, C., Small, M.J. and Wu, F. (2000). An integrated risk model of a drinking-water - borne cryptosporidiosis outbreak. Risk Anal., 20(4): 495-511.
Diallo, M.B.C., Anceno, A.J., Tawatsupa, B., Houpt, E.R., Wangsuphachart, V. and Shipin, O.V. (2008). Infection risk assessment of diarrhea-related pathogens in a tropical canal network. Sci. Total Environ., 407(1): 223-232.
Emelko, M.B., Schmidt, P.J. and Roberson, J.A. (2008). Quantification of uncertainty in microbial data - reporting and regulatory implications. J. Am. Water Works Assoc., 100(3): 94-104+14.
Gibney, K.B., O'Toole, J., Sinclair, M. and Leder, K. (2014). Disease burden of selected gastrointestinal pathogens in australia, 2010. Int. J. Infect. Dis., 28: e176-e185.
Haas, C.N., Rose, J.B. and Gerba, C.P. (1999). Quantitative microbial risk assessment. Quantitative microbial risk assessment. John Wiley & Sons, Inc, New York, New York.
Havelaar, A.H. and Melse, J.M. (2003). Quantifying public health risk in the WHO Guidelines for drinking-water quality: a burden of disease approach. Rijkinstituut voor Volskgezondheid en Milieu, RIVM 734301022, Bilthoven, The Netherlands.
Health Canada (2011). Guidelines for Canadian drinking water quality: Guideline technical document - enteric viruses. Water, Air and Climate Change Bureau, Healthy Environments and Consumer Safety Branch, Health Canada, Ottawa, Ontario. Available at: www.canada.ca/en/health-canada/services/publications/healthy-living/guidelines-canadian-drinking-water-quality-guideline-technical-document-enteric-viruses.html
Health Canada (2012). Guidelines for Canadian drinking water quality: Guideline technical document - enteric protozoa: Giardia and Cryptosporidium. Water, Air and Climate Change Bureau, Healthy Environments and Consumer Safety Branch, Health Canada, Ottawa, Ontario. Available at: www.canada.ca/en/health-canada/services/environmental-workplace-health/reports-publications/water-quality/enteric-protozoa-giardia-cryptosporidium.html.
Health Canada (2013). Guidance on the use of the microbiological drinking water quality guidelines. Water and Air Quality Bureau, Healthy Environments and Consumer Safety Branch, Health Canada, Ottawa, Ontario. Available at: www.canada.ca/en/health-canada/services/environmental-workplace-health/reports-publications/water-quality/guidance-use-microbiological-drinking-water-quality-guidelines-health-canada-2013.html
Hijnen, W.A.M. and Medema, G. (2007). Elimination of micro-organisms by drinking water treatment process: A review. Kiwa Water Research, Nieuwegein, The Netherlands.
Jaidi, K., Barbeau, B., Carrière, A., Desjardins, R. and Prévost, M. (2009). Including operational data in QMRA model: Development and impact of model inputs. J. Water Health, 7(1): 77-95.
Macler, B.A. and Regli, S. (1993). Use of microbial risk assessment in setting US drinking water standards. Int. J. Food Microbiol., 18(4): 245-256.
Makri, A., Modarres, R. and Parkin, R. (2004). Cryptosporidiosis susceptibility and risk: A case study. Risk Anal., 24(1): 209-220.
Mara, D.D., Sleigh, P.A., Blumenthal, U.J. and Carr, R.M. (2007). Health risks in wastewater irrigation: Comparing estimates from quantitative microbial risk analyses and epidemiological studies. J. Water Health, 5(1): 39-50.
Martins, M.T., Rivera, I.G., Clark, D.L. and Olson, B.H. (1992). Detection of virulence factors in culturable escherichia coli isolates from water samples by DNA probes and recovery of toxin-bearing strains in minimal o-nitrophenol-ß-D-galactopyranoside-4-methylumbelliferyl-ß-D- glucuronide media. Appl. Environ. Microbiol., 58(9): 3095-3100.
Medema, G., Teunis, P., Blokker, M., Deere, D., Davison, A., Charles, P., and Loret, J-F. (2009). Risk assessment of cryptosporidium in drinking water. WHO Press, World Health Organization, Geneva, Switzerland.
Messner, M.J., Chappell, C.L. and Okhuysen, P.C. (2001). Risk assessment for cryptosporidium: A hierarchical bayesian analysis of human dose response data. Water Res., 35(16): 3934-3940.
MOE (2006). Procedure for disinfection of drinking water in ontario. as adopted by reference by ontario regulation 170/03 under the safe drinking water act. Ontario Ministry of the Environment, PIBS 4448e01, Toronto, Ontario.
Murray, C. and Lopez, A. (1996). Global health statistics. Harvard School of Public Health, Cambridge, Massachusetts.
Murray, C.J.L. and Lopez, A.D. (1996). The global burden of disease: A comprehensive assessment of mortality and disability from disease, injury and risk factors in 1990 and projected to 2020. Harvard University Press, Cambridge, Massachusetts.
Nash, T.E., Herrington, D.A., Losonsky, G.A. and Levine, M.M. (1987). Experimental human infections with giardia lamblia. J. Infect. Dis., 156(6): 974-984.
Ongerth, J.E. (2013). ICR SS protozoan data site-by-site: A picture of cryptosporidium and giardia in U.S. surface water. Environ. Sci. Technol., 47(18): 10145-10154.
Pouillot, R., Beaudeau, P., Denis, J.-. and Derouin, F. (2004). A quantitative risk assessment of waterborne cryptosporidiosis in france using second-order monte carlo simulation. Risk Anal., 24(1): 1-17.
Regli, S., Rose, J.B., Haas, C.N. and Gerba, C.P. (1991). Modeling the risk from giardia and viruses in drinking water. J. Am. Water Works Assoc., 83(11): 76-84.
Rose, J.B. and Gerba, C.P. (1991). Use of risk assessment for development of microbial standards. Water Sci. Technol., 24(2): 29-34.
Schijven, J., Derx, J., de Roda Husman, A.M., Blaschke, A.P. and Farnleitner, A.H. (2015). QMRAcatch: Microbial quality simulation of water resources including infection risk assessment. J. Environ. Qual., 44(5): 1491-1502.
Schijven, J.F., Teunis, P.F.M., Rutjes, S.A., Bouwknegt, M. and de Roda Husman, A.M. (2011). QMRAspot: A tool for quantitative microbial risk assessment from surface water to potable water. Water Res., 45(17): 5564-5576.
Signor, R.S. and Ashbolt, N.J. (2006). Pathogen monitoring offers questionable protection against drinking-water risks: A QMRA (quantitative microbial risk analysis) approach to assess management strategies. pp. 261-268. Available at: www.scopus.com/inward/record.url?eid=2-s2.0-33749155605&partnerID=40&md5=043744c9cf5e3b589eca03b2ba9e8313.
Signor, R.S. and Ashbolt, N.J. (2009). Comparing probabilistic microbial risk assessments for drinking water against daily rather than annualised infection probability targets. J. Water Health, 7(4): 535-543.
Smeets, P.W.M.H., van der Helm, A.W.C., Dullemont, Y.J., Rietveld, L.C., van Dijk, J.C. and Medema, G.J. (2006). Inactivation of escherichia coli by ozone under bench-scale plug flow and full-scale hydraulic conditions. Water Res., 40(17): 3239-3248.
Smeets, P.W.M.H., van Dijk, J.C., Stanfield, G., Rietveld, L.C. and Medema, G.J. (2007). How can the UK statutory cryptosporidium monitoring be used for quantitative risk assessment of cryptosporidium in drinking water? J. Water Health, 5(SUPPL. 1): 107-118.
Smeets, P.W.M.H., Dullemont, Y.J., Van Gelder, P.H.A.J.M., Van Dijk, J.C. and Medema, G.J. (2008). Improved methods for modelling drinking water treatment in quantitative microbial risk assessment; a case study of campylobacter reduction by filtration and ozonation. J. Water Health, 6(3): 301-314.
Soller, J.A. and Eisenberg, J.N.S. (2008). An evaluation of parsimony for microbial risk assessment models. Environmetrics, 19(1): 61-78.
Statistics Canada (2004). Data source: Canadian community health survey, cycle 2.2 - nutrition (wave 3): General health and 24-hour dietary recall. (Share File). Statistics Canada, Ottawa, Ontario.
Statistics Canada (2008). User guide: Canadian community health survey (CCHS), cycle 2.2 (2004), nutrition - general health (including vitamin & mineral supplements) & 24-hour dietary recall components. Statistics Canada, Ottawa, Ontario.
Statistics Canada (2011). Census data navigator - age and sex for Canada. Available at: www12.statcan.gc.ca/census-recensement/2011/dp-pd/map-carte/index-eng.cfm.
Statistics Canada (2012). Life table, canada, provinces and territories 2007-2009. Ministry of Industry, ISBN: 978-1-100-21498-6.
Strachan, N.J.C., Doyle, M.P., Kasuga, F., Rotariu, O. and Ogden, I.D. (2005). Dose response modelling of escherichia coli O157 incorporating data from foodborne and environmental outbreaks. Int. J. Food Microbiol., 103(1): 35-47.
Teunis, P.F.M., Medema, G.J., Kruidenier, L. and Havelaar, A.H. (1997). Assessment of the risk of infection by cryptosporidium or giardia in drinking water from a surface water source. Water Res., 31(6): 1333-1346.
Teunis, P.F.M., Moe, C.L., Liu, P., Miller, S.E., Lindesmith, L., Baric, R.S., Le Pendu, J. and Calderon, R.L. (2008). Norwalk virus: How infectious is it? J. Med. Virol., 80(8): 1468-1476.
Teunis, P.F.M., Rutjes, S.A., Westrell, T. and de Roda Husman, A.M. (2009). Characterization of drinking water treatment for virus risk assessment. Water Res., 43(2): 395-404.
U.S. EPA (2003). LT1ESWTR disinfection profiling and benchmarking technical guidance manual. Office of Water, U.S. Environmental Protection Agency, Washington, DC.
U.S. EPA (2005). Economic analysis for the final Long Term 2 Enhanced Surface Water Treatment Rule. U.S. Environmental Protection Agency, EPA 815-R-06-001.
U.S. EPA (2006). Economic analysis for the final Groundwater Rule. U.S. Environmental Protection Agency, EPA 815-R-06-014.
U.S. EPA (2014). Microbiological risk assessment (MRA) tools, methods, and approaches for water media. Office of Water, Office of Science and Technology. U.S. Environmental Protection Agency, EPA-820-R-14-009, Washington, DC.
WHO (2016). Quantitative microbial risk assessment: Application for water safety management. World Health Organization, Geneva, Switzerland.
C.2 Acronyms
- CSTR
- continuously stirred tank reactor
- CT
- concentration × time
- DALYs
- disability adjusted life years
- EPA
- Environmental Protection Agency (U.S.)
- ESRD
- end-stage renal disease
- GBS
- Guillain-Barré syndrome
- HC
- QMRA Health Canada quantitative microbial risk assessment (model)
- HUS
- hemolytic uremic syndrome
- LOD
- limit of detection
- LYL
- life years lost
- QMRA
- quantitative microbial risk assessment
- UV
- ultraviolet
- WHO
- World Health Organization
- YLD
- years lived with a disability
- UV
- ultraviolet