Archived - Shiga toxin-producing Escherichia coli, British Columbia 2011–2017

 Download this article as a PDF
Download this article as a PDFPublished by: The Public Health Agency of Canada
Issue: Volume 45–9: Outbreaks
Date published: September 5, 2019
ISSN: 1481-8531
Submit a manuscript
About CCDR
Browse
Volume 45–9, September 5, 2019: Outbreaks
Surveillance
Shiga toxin–producing Escherichia coli in British Columbia, 2011–2017: Analysis to inform exclusion guidelines
K Noftall1,2, M Taylor1, L Hoang3,4, E Galanis1,2
Affiliations
1 BC Centre for Disease Control, Vancouver, BC
2 School of Population and Public Health, University of British Columbia, Vancouver, BC
3 BC Centre for Disease Control Public Health Laboratory, Vancouver, BC
4 Department of Pathology and Laboratory Medicine, Faculty of Medicine, University of British Columbia, Vancouver, BC
Correspondence
Suggested citation
Noftall K, Taylor M, Hoang L, Galanis E. Shiga toxin–producing Escherichia coli in British Columbia, 2011–2017: Analysis to inform exclusion guidelines. Can Commun Dis Rep 2019;45(9):238–43. https://doi.org/10.14745/ccdr.v45i09a03
Keywords: Shiga toxin, Escherichia coli, guidelines, surveillance, STEC
Abstract
Background: Shiga toxin–producing Escherichia coli (STEC) can cause severe illness including bloody diarrhea and hemolytic-uremic syndrome (HUS) through the production of Shiga toxins 1 (Stx1) and 2 (Stx2). E. coli O157:H7 was the most common serotype detected in the 1980s to 1990s, but improvements in laboratory methods have led to increased detection of non-O157 STEC. Non-O157 STEC producing only Stx1 tend to cause milder clinical illness. Exclusion guidelines restrict return to high-risk work or settings for STEC cases, but most do not differentiate between STEC serogroups and Stx type.
Objective: To analyze British Columbia (BC) laboratory and surveillance data to inform the BC STEC exclusion guideline.
Methods: For all STEC cases reported in BC in 2011–2017, laboratory and epidemiological data were obtained through provincial laboratory and reportable disease electronic systems, respectively. Incidence was measured for all STEC combined as well as by serogroup. Associations were measured between serogroups, Stx types and clinical outcomes.
Results: Over the seven year period, 984 cases of STEC were reported. A decrease in O157 incidence was observed, while non-O157 rates increased. The O157 serogroup was significantly associated with Stx2. Significant associations were observed between Stx2 and bloody diarrhea, hospitalization and HUS.
Conclusion: The epidemiology of STEC has changed in BC as laboratories increasingly distinguish between O157 and non-O157 cases and identify Stx type. It appears that non-O157 cases with Stx1 are less severe than O157 cases with Stx2. The BC STEC exclusion guidelines were updated as a result of this analysis.
Introduction
Escherichia coli has long been known to be part of the normal flora of the gastrointestinal tract. Some E. coli strains evolved to cause human illness by acquiring virulence factors such as Shiga toxin. These organisms have been renamed Shiga toxin–producing E. coli (STEC)Footnote 1. The best known STEC serotype, E. coli O157:H7, was first detected in the 1980s in patients with serious illnesses including hemorrhagic colitis and hemolytic-uremic syndrome (HUS)Footnote 2Footnote 3. In the 1990s, once laboratories started to routinely test for E. coli O157:H7, it became recognized as the cause of large and serious outbreaks mainly associated with undercooked beefFootnote 4Footnote 5.
In the early 2000s, laboratories began to implement methods to detect other STEC serotypesFootnote 6Footnote 7. This led to the recognition that non-O157 STEC can contaminate a wide variety of sources and cause large outbreaksFootnote 8Footnote 9. However, non‑O157 STEC remain underreported because many frontline laboratories do not yet routinely test for themFootnote 10. Nonetheless, by 2016, approximately 35% of all STEC infections reported to the Canadian National Enteric Surveillance Program were caused by non-O157 serotypesFootnote 11.
STEC stx genes produce Shiga toxins 1 (Stx1) and 2 (Stx2). Stx2 is associated with severe illness and HUSFootnote 8Footnote 12Footnote 13Footnote 14Footnote 15Footnote 16Footnote 17Footnote 18Footnote 19Footnote 20. Furthermore, Stx2 is associated with STEC O157, which explains this serogroup’s higher pathogenicityFootnote 17Footnote 21Footnote 22. A polymerase chain reaction (PCR) test was implemented at the British Columbia Centre for Disease Control Public Health Laboratory (BCCDC PHL) in 2013 to detect and differentiate between stx 1 and 2 genes.
In order to protect the population from severe infection, most public health authorities exclude people infected with STEC from working or attending certain high-risk settingsFootnote 23Footnote 24. For example, the 2006 BCCDC guidelines stated that a person diagnosed with STEC who works as a food handler, health care worker or child care worker or who attends a child care facility should be excluded from that setting until they have provided two negative stool samplesFootnote 25.
The increasing detection of non-O157 STEC and their association with Stx1 and less severe disease led us to question the appropriateness of exclusion guidelines that do not differentiate between serogroups and Stx types. A review of existing guidelines demonstrated that four regions had recently updated their recommendations to stratify exclusion from workplaces and daycares by serogroup and Stx typeFootnote 26Footnote 27Footnote 28Footnote 29.
The purpose of this work was to analyze British Columbia (BC) laboratory and surveillance data to inform the BC STEC exclusion guideline.
Methods
Study sample
This study included all confirmed cases of STEC who were BC residents and reported in the provincial electronic public health information system between January 1, 2011 and December 31, 2017Footnote 30. For these cases, all STEC culture, serogroup and PCR stx results were accessed from the BCCDC PHL information system.
Laboratory data
In BC, stool specimens may be processed at a private laboratory, hospital laboratory or the BCCDC PHL. At private and hospital laboratories, STEC are identified using culture or molecular methods. STEC isolates that are Stx positive or O157, as well as specimens that are visibly bloody or that came from a case diagnosed with HUS, are sent to the BCCDC PHL for further testing and serotyping. Until 2013, Stx was detected using a Vero cell assay at the BCCDC PHL; since 2013, stx 1 and 2 genes have been detected by PCR from stool samples and from suspected STEC isolates recovered from stx-positive stool samples from private and hospital laboratories. Positive stx isolates are serotyped for O157 antigen, H7 antigen and an O-Typer PCR covering the six most common serogroups seen in BC: O26, O45, O103, O111, O121 and O145. Any stx-positive isolate that cannot be serotyped is sent to the National Microbiology Laboratory for identification.
Variables included from BCCDC PHL data were Stx type and serogroup. If Vero cell assay was used, the Stx result was reported as positive; when PCR was performed, the stx gene was recorded. When multiple laboratory test results were available for individual cases, the most specific serotype result was included.
Surveillance data
STEC cases are reportable in BC and are interviewed by public health professionals using a standard surveillance formFootnote 30Footnote 31. All health authorities report STEC cases provincially through the BC provincial electronic public health information system. Self-reported data on demographics, clinical symptoms, hospitalization and HUS status were extracted from this system. Laboratory data may also be entered into this system as part of routine surveillance by the health authority.
Hospitalization status and exclusion information were only available for cases from 2015–2017. Clinical data were available from four of five health authorities. Cases were considered to have a symptom or outcome if “yes” was recorded for that data field. If no laboratory results were available from BCCDC PHL, the laboratory information reported by the health authorities into the public health information system was used.
Analysis
BCCDC STEC surveillance data and BCCDC PHL STEC lab data from 2011 to 2017 were linked together by a common case identifier. Incidence rates were calculated using the BC population dataFootnote 32. Differences in STEC incidence rates between females and males were conducted using the Wilcoxon rank sum test. Clinical information was compared by serogroups and by Stx type for all symptoms as well as for HUS and hospitalization using Fisher exact test. An alpha level of 0.05 was used for significance, and all statistical tests were conducted using R statistical computing programFootnote 33. When exclusion information was available, excluded cases were categorized by serogroup and Stx type.
Results
Between 2011 and 2017, 984 cases of STEC were reported in BC, with an average of 141 cases per year (range: 108–184). The overall STEC incidence during this time period was 3.0 cases per 100,000. Outbreaks occurred in 2013, 2016 and 2017. There were more female (57.4%) than male (42.6%) cases, but there was no significant difference in rates by sex for all STEC cases or by serogroups (data not shown). The median age was 31 years (range <1–113), and the highest incidence was in 1–9 year olds at 4.9 cases per 100,000.
In total, 58 serogroups were identified. Nearly 10% of samples were culture negative and stx positive only (Table 1). The six most common serogroups were O157, O121, O26, O103, O117 and O111. The most reported serogroup was O157 for every year except 2017, when O121 cases were more common.
| Serogroup | N | (%) |
|---|---|---|
| O157 | 369 | (37.5) |
| O121 | 102 | (10.4) |
| O26 | 86 | (8.7) |
| O103 | 44 | (4.5) |
| O117 | 38 | (3.9) |
| O111 | 34 | (3.5) |
| Other serogroups | 152 | (15.4) |
| Stx positive only | 93 | (9.5) |
| UnknownFootnote a of Table 1 | 66 | (6.7) |
| Shiga toxin | N | (%) |
| Stx positiveFootnote b of Table 1 | 111 | (11.3) |
| Stx1 | 270 | (27.4) |
| Stx1 and Stx2 | 239 | (24.3) |
| Stx2 | 274 | (27.8) |
| Stx unknownFootnote c of Table 1 | 90 | (9.1) |
Overall, O157 rates have declined from 1.1 to 0.5 per 100,000 from 2011 to 2017 (Figure 1). Non-O157 rates have increased greatly from 0.6 to 2.4 per 100,000 between 2011 and 2017.
Figure 1: Shiga toxin–producing Escherichia coli incidence by serogroup, British Columbia, 2011–2017 (N=984)
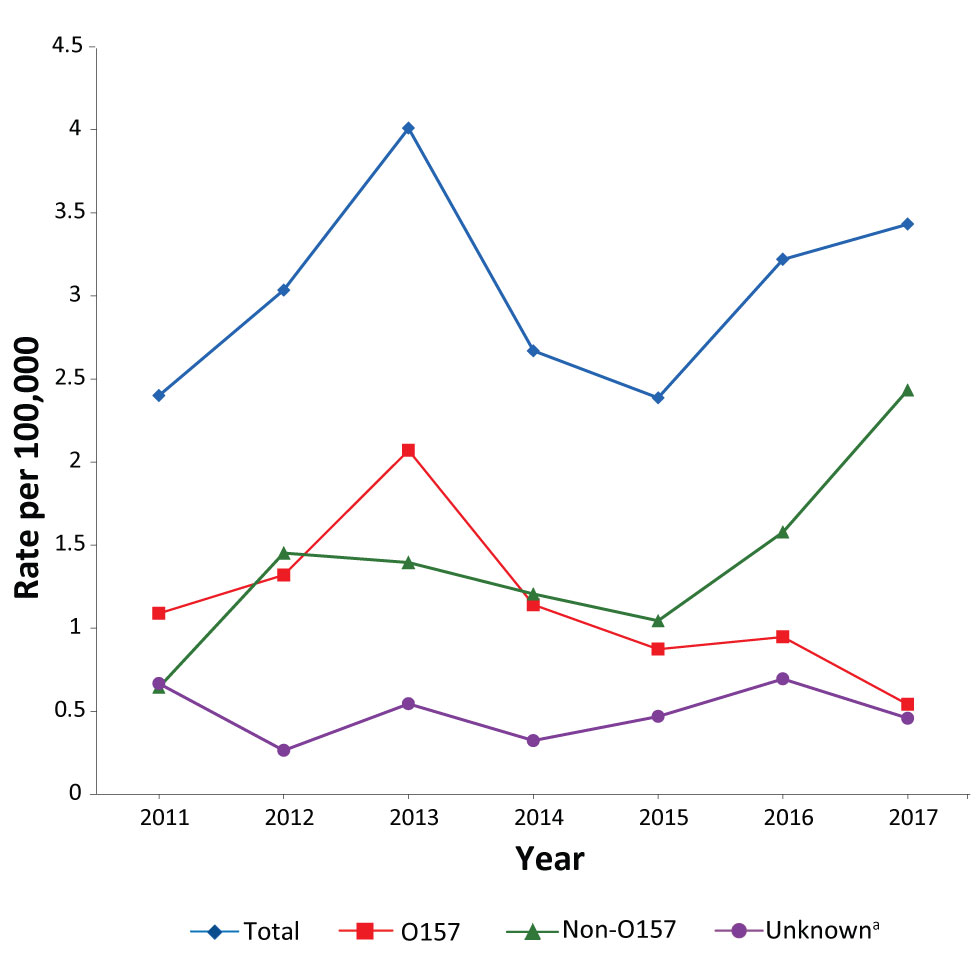
Text description: Figure 1
Figure 1: Shiga toxin–producing Escherichia coli incidence by serogroup, British Columbia, 2011–2017 (N=984)
| Year | O157 | Non-O157 | UnknownFigure 1 footnote a | Total |
|---|---|---|---|---|
| 2011 | 1.1 | 0.6 | 0.7 | 2.4 |
| 2012 | 1.3 | 1.5 | 0.3 | 3.0 |
| 2013 | 2.1 | 1.4 | 0.5 | 4.0 |
| 2014 | 1.1 | 1.2 | 0.3 | 2.7 |
| 2015 | 0.9 | 1.0 | 0.5 | 2.4 |
| 2016 | 0.9 | 1.6 | 0.7 | 3.2 |
| 2017 | 0.5 | 2.4 | 0.5 | 3.4 |
The presence of Stx was confirmed in 894 cases (90.1%; Table 1). For 111 cases (11.3%), Stx was detected by Vero cell assay only, and specific Stx type was not available. Of the 783 cases where the stx gene was identified by PCR, 270 (34.5%) were positive for stx1 only, 274 (35.0%) were positive for stx2 only, and 239 (30.5%) were from STEC that contained both stx1 and stx2. Therefore, 65.5% of cases where the stx gene was detected tested positive for stx2 (alone or in combination). When both serogroup and stx gene were identified (N=711), 97.7% of O157 STEC contained stx2 alone or in combination compared to 42.9% of non-O157 serogroups [odd ratio (OR) = 56.5; p≤0.001].
Clinical information was available for 594 cases (60.4%). The most commonly reported symptoms were diarrhea (90.2%), abdominal discomfort (70.2%), bloody diarrhea (56.6%), vomiting (24.6%), fever (21.4%) and fatigue (13.6%). Eighteen cases (3.0%) were recorded as having HUS at the time of the public health interview. Cases with E. coli O157 infection were more likely to have bloody diarrhea, fatigue and hospitalization than non-O157 cases (Table 2). Although a larger proportion of O157 cases (4.9%) had HUS than non-O157 cases (1.9%), this was not statistically significant. Cases with stx2 were significantly more likely to have bloody diarrhea and HUS and be admitted to hospital than cases with stx1 only (Table 2).
| Clinical outcome | Serogroup | Shiga toxin type | ||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| O157 | Non-O157 | p-value | Stx2 present | Stx1 only | p-value | |||||
| N | (%) | N | (%) | N | (%) | N | (%) | |||
| Abdominal discomfort | 177 | (72.5) | 189 | (70.3) | 0.625 | 262 | (73.8) | 106 | (69.3) | 0.330 |
| Diarrhea | 221 | (90.5) | 250 | (92.9) | 0.339 | 321 | (90.4) | 138 | (90.2) | 1.000 |
| Bloody diarrhea | 168 | (68.9) | 136 | (50.6) | <0.001 | 231 | (65.1) | 69 | (45.1) | <0.001 |
| Fatigue | 46 | (18.9) | 25 | (9.3) | 0.002 | 53 | (14.9) | 17 | (11.1) | 0.326 |
| Fever | 57 | (23.4) | 50 | (18.6) | 0.193 | 78 | (22.0) | 27 | (17.6) | 0.285 |
| Vomiting | 63 | (25.8) | 64 | (23.8) | 0.610 | 85 | (23.9) | 34 | (22.2) | 0.733 |
| HUS | 12 | (4.9) | 5 | (1.9) | 0.081 | 15 | (4.2) | 1 | (0.7) | 0.048 |
| HospitalisedFootnote b of Table 2 | 29 | (35.4) | 25 | (16.1) | 0.001 | 52 | (31.7) | 13 | (13.8) | 0.002 |
Exclusion information was available for 276 (64%) cases between 2015 and 2017. Forty-three cases (16%) had been excluded from high-risk occupations or settings. Ten of the cases were positive for O157 (23.3%), 28 were positive for a non-O157 STEC (65.1%) and five had an unknown serogroup (11.6%). Twenty-one cases (48.8%) had STEC with stx2 present, 20 (46.5%) had STEC with stx1 only and two (4.7%) had STEC with indeterminate stx PCR results. All O157 STEC cases were stx2 positive.
Discussion
Analysis of laboratory and surveillance data in BC showed a shifting pattern in the distribution of STEC serogroups between 2011 and 2017 with the incidence of O157 declining and the incidence of non-O157 increasing, particularly between 2015 and 2017. This is consistent with the decline in the incidence of O157 that has been observed across CanadaFootnote 34 and the United StatesFootnote 6Footnote 8Footnote 22Footnote 35Footnote 36. The decrease in O157 in Canada may be due to sanitary improvements in beef processing as well as better food safety education to consumersFootnote 34. The increase in incidence of non-O157 in BC and elsewhere may be due to the increase in use of laboratory methods enabling the detection of non-O157 strainsFootnote 6Footnote 36.
A strong association was seen between the O157 serogroup and Stx2 with nearly 98% of O157 cases being positive for Stx2, consistent with earlier studiesFootnote 15Footnote 16Footnote 21Footnote 22. Also observed were the associations between O157 STEC cases and severe clinical outcomes such as hemorrhagic colitis and hospitalizationFootnote 17Footnote 22Footnote 37. There was a significant relationship between stx2 and HUS, bloody diarrhea and hospitalization. Stx variants (e.g. stx2d), which have been shown to be more specific predictors of severe illness than Stx type, are not yet available in BCFootnote 13Footnote 15Footnote 16Footnote 18Footnote 19Footnote 20Footnote 38.
This analysis was conducted to inform the BC STEC exclusion guideline. Consistent with our findings and other recently updated guidelines stratifying exclusion by serogroup and StxFootnote 26Footnote 27Footnote 28Footnote 29, the new BC guideline now reflects the evidence that STEC producing Stx1 only leads to less severe illnessFootnote 39. Based on the revised guideline whereby non-O157 stx1 only cases without severe clinical symptoms could return to high-risk work or settings when symptom-free, 16 excluded cases (37.2%) between 2015 and 2017 would have had their exclusion lifted earlier. Assuming the median duration of symptoms is seven days and the median duration of shedding is 20 daysFootnote 40Footnote 41, this could have resulted in avoiding up to 208 days of lost productivity for these individuals.
This study has a few limitations. Laboratory practice for Stx detection changed during the study period, so we were unable to differentiate Stx types between 2011 and 2013. Symptoms were self-reported and only available for four of the five health authorities in the province. Despite this, the proportion of cases with symptoms was similar to that observed in other studiesFootnote 22Footnote 42. There is no reason to believe symptoms and outcomes for the same disease would be different for the one health authority from which these data were unavailable. The number of HUS cases was likely an underestimate given that HUS may not have developed at the time of the public health interview, which occurs one to two weeks after symptom onsetFootnote 43. Nonetheless, a significant association with stx2 was observed, and this association may be even stronger with a higher number of HUS cases. Exclusion information was not available prior to 2015 and was not always recorded after that. Because of this, only a descriptive analysis was performed for excluded cases.
An evaluation of this guideline change in BC is recommended to ensure its impact was as initially intended. Furthermore, the guideline will need to be reviewed as laboratory methods for STEC and Stx detection progress, such as by testing for specific Stx variants.
Conclusion
Data from BC is consistent with observations elsewhere that STEC bacteria cause a spectrum of illness and this is, at least in part, determined by Stx type. As a result of this analysis, the BC STEC exclusion guideline was updated in February 2019 to allow cases infected with non-O157 stx1 only STEC without severe clinical symptoms to return to high-risk settings when symptom-free.
Authors’ Statement
- KN — Analyzed the data and drafted the paper
- MT — Conceptualized the study, interpreted the data and revised the paper
- LH — Interpreted the data and revised the paper
- EG — Conceptualized the study, interpreted the data, drafted certain sections and revised the paper
Conflict of interest
None.
Acknowledgements
The authors are grateful to A Paccagnella and British Columbia Enteric Policy Group members (O Bitzikos, D Hoyano, M McKinley, S Mema, P Rydings, J Stone, D Tam) who contributed data or participated in the decision making about the ensuing guideline changes. The authors also want to acknowledge the editorial contributions of the Editor-in-Chief of the Canada Communicable Disease Report (CCDR).
